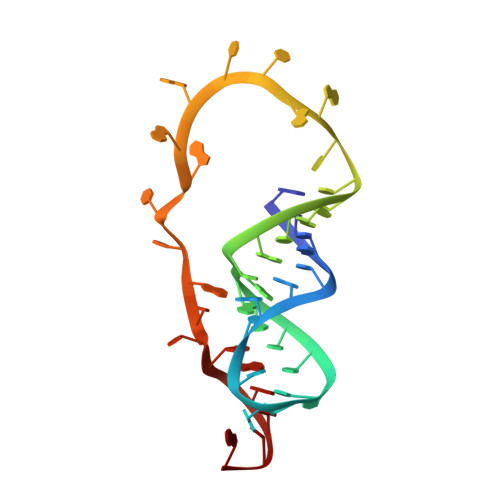
The structure of the SAM/SAH-binding riboswitch.
Weickhmann, A.K., Keller, H., Wurm, J.P., Strebitzer, E., Juen, M.A., Kremser, J., Weinberg, Z., Kreutz, C., Duchardt-Ferner, E., Wohnert, J.(2019) Nucleic Acids Res 47: 2654-2665
- PubMed: 30590743 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gky1283
- Primary Citation Related Structures:
6HAG - PubMed Abstract:
S-adenosylmethionine (SAM) is a central metabolite since it is used as a methyl group donor in many different biochemical reactions. Many bacteria control intracellular SAM concentrations using riboswitch-based mechanisms. A number of structurally different riboswitch families specifically bind to SAM and mainly regulate the transcription or the translation of SAM-biosynthetic enzymes. In addition, a highly specific riboswitch class recognizes S-adenosylhomocysteine (SAH)-the product of SAM-dependent methyl group transfer reactions-and regulates enzymes responsible for SAH hydrolysis. High-resolution structures are available for many of these riboswitch classes and illustrate how they discriminate between the two structurally similar ligands SAM and SAH. The so-called SAM/SAH riboswitch class binds both ligands with similar affinities and is structurally not yet characterized. Here, we present a high-resolution nuclear magnetic resonance structure of a member of the SAM/SAH-riboswitch class in complex with SAH. Ligand binding induces pseudoknot formation and sequestration of the ribosome binding site. Thus, the SAM/SAH-riboswitches are translational 'OFF'-switches. Our results establish a structural basis for the unusual bispecificity of this riboswitch class. In conjunction with genomic data our structure suggests that the SAM/SAH-riboswitches might be an evolutionary late invention and not a remnant of a primordial RNA-world as suggested for other riboswitches.
- Institute for Molecular Biosciences and Center for Biomolecular Magnetic Resonance (BMRZ), Goethe-University Frankfurt, Max-von-Laue-Strasse 9, 60438 Frankfurt/M., Germany.
Organizational Affiliation: