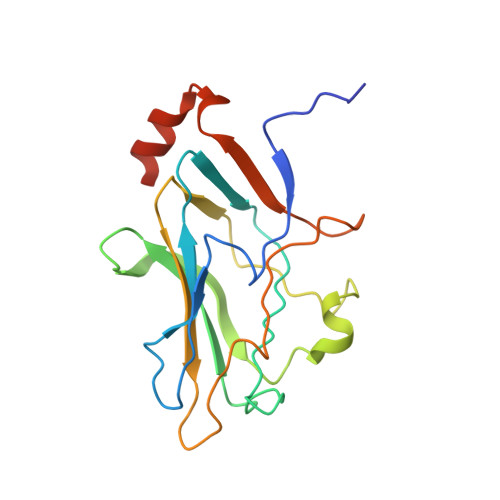
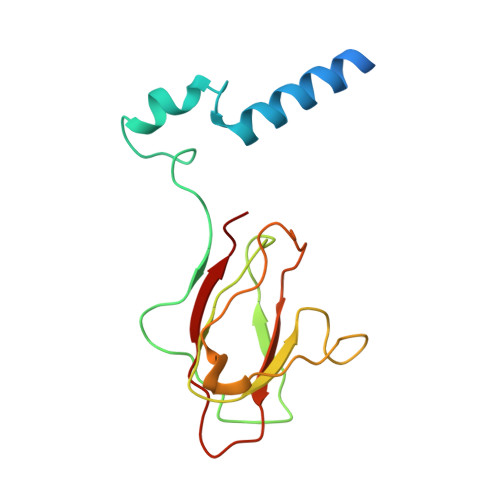
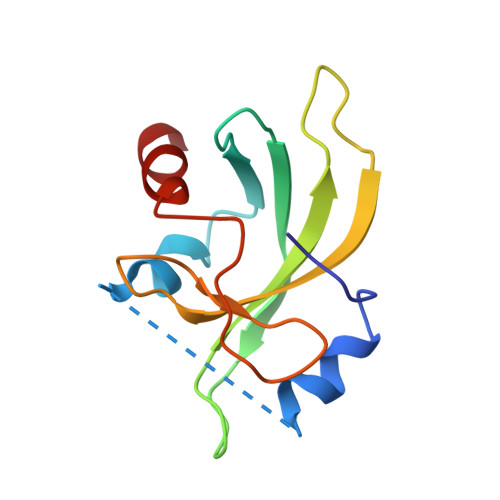
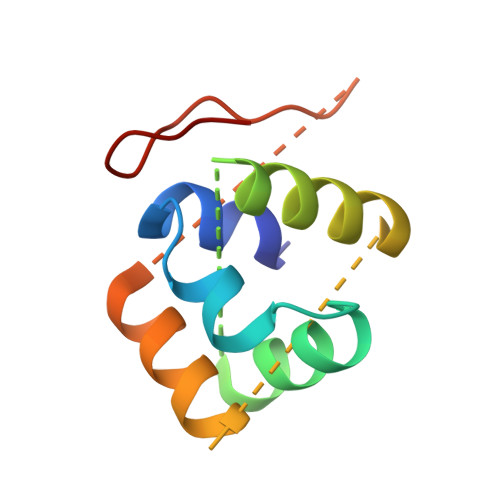
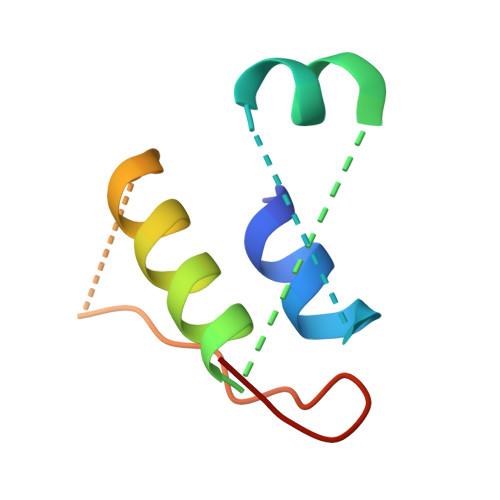
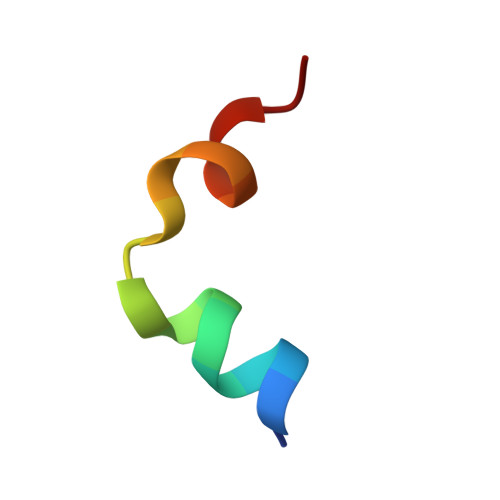
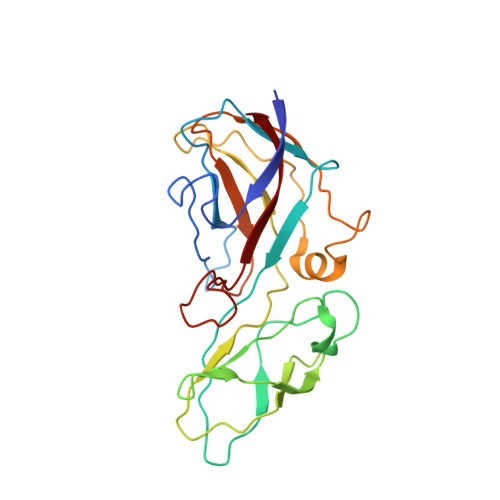
Structural basis for assembly of vertical single beta-barrel viruses.
Santos-Perez, I., Charro, D., Gil-Carton, D., Azkargorta, M., Elortza, F., Bamford, D.H., Oksanen, H.M., Abrescia, N.G.A.(2019) Nat Commun 10: 1184-1184
- PubMed: 30862777 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-08927-2
- Primary Citation Related Structures:
6H82, 6H9C - PubMed Abstract:
The vertical double β-barrel major capsid protein (MCP) fold, fingerprint of the PRD1-adeno viral lineage, is widespread in many viruses infecting organisms across the three domains of life. The discovery of PRD1-like viruses with two MCPs challenged the known assembly principles. Here, we present the cryo-electron microscopy (cryo-EM) structures of the archaeal, halophilic, internal membrane-containing Haloarcula californiae icosahedral virus 1 (HCIV-1) and Haloarcula hispanica icosahedral virus 2 (HHIV-2) at 3.7 and 3.8 Å resolution, respectively. Our structures reveal proteins located beneath the morphologically distinct two- and three-tower capsomers and homopentameric membrane proteins at the vertices that orchestrate the positioning of pre-formed vertical single β-barrel MCP heterodimers. The cryo-EM based structures together with the proteomics data provide insights into the assembly mechanism of this type of viruses and into those with membrane-less double β-barrel MCPs.
- Molecular Recognition and Host-pathogen Interactions Programme, CIC bioGUNE, CIBERehd, Bizkaia Technology Park, 48160, Derio, Spain.
Organizational Affiliation: