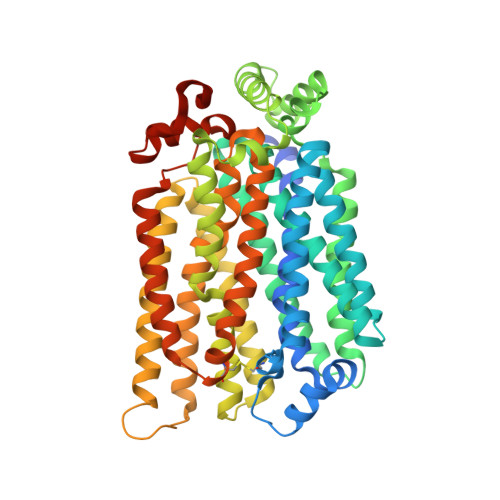
Crystal structure of the plant symporter STP10 illuminates sugar uptake mechanism in monosaccharide transporter superfamily.
Paulsen, P.A., Custodio, T.F., Pedersen, B.P.(2019) Nat Commun 10: 407-407
- PubMed: 30679446 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-08176-9
- Primary Citation Related Structures:
6H7D - PubMed Abstract:
Plants are dependent on controlled sugar uptake for correct organ development and sugar storage, and apoplastic sugar depletion is a defense strategy against microbial infections like rust and mildew. Uptake of glucose and other monosaccharides is mediated by Sugar Transport Proteins, proton-coupled symporters from the Monosaccharide Transporter (MST) superfamily. We present the 2.4 Å structure of Arabidopsis thaliana high affinity sugar transport protein, STP10, with glucose bound. The structure explains high affinity sugar recognition and suggests a proton donor/acceptor pair that links sugar transport to proton translocation. It contains a Lid domain, conserved in all STPs, that locks the mobile transmembrane domains through a disulfide bridge, and creates a protected environment which allows efficient coupling of the proton gradient to drive sugar uptake. The STP10 structure illuminates fundamental principles of sugar transport in the MST superfamily with implications for both plant antimicrobial defense, organ development and sugar storage.
- Department of Molecular Biology and Genetics, Aarhus University, Gustav Wieds Vej 10, DK-8000, Aarhus C, Denmark.
Organizational Affiliation: