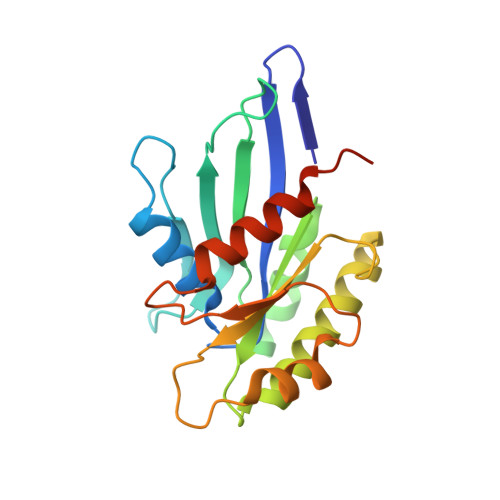
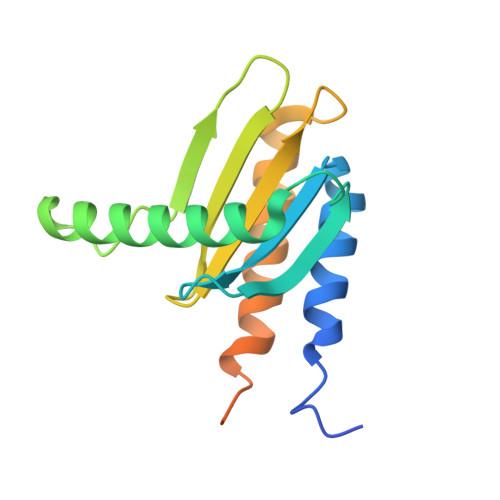
MglA functions as a three-state GTPase to control movement reversals of Myxococcus xanthus.
Galicia, C., Lhospice, S., Varela, P.F., Trapani, S., Zhang, W., Navaza, J., Herrou, J., Mignot, T., Cherfils, J.(2019) Nat Commun 10: 5300-5300
- PubMed: 31757955 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-13274-3
- Primary Citation Related Structures:
6H17, 6H35, 6H5B, 6HJH, 6HJM, 6HJO - PubMed Abstract:
In Myxococcus xanthus, directed movement is controlled by pole-to-pole oscillations of the small GTPase MglA and its GAP MglB. Direction reversals require that MglA is inactivated by MglB, yet paradoxically MglA and MglB are located at opposite poles at reversal initiation. Here we report the complete MglA/MglB structural cycle combined to GAP kinetics and in vivo motility assays, which uncovers that MglA is a three-state GTPase and suggests a molecular mechanism for concerted MglA/MglB relocalizations. We show that MglA has an atypical GTP-bound state (MglA-GTP*) that is refractory to MglB and is re-sensitized by a feedback mechanism operated by MglA-GDP. By identifying and mutating the pole-binding region of MglB, we then provide evidence that the MglA-GTP* state exists in vivo. These data support a model in which MglA-GDP acts as a soluble messenger to convert polar MglA-GTP* into a diffusible MglA-GTP species that re-localizes to the opposite pole during reversals.
- Laboratoire de Biologie et Pharmacologie Appliquée, CNRS and Ecole Normale Supérieure Paris-Saclay, Cachan, France.
Organizational Affiliation: