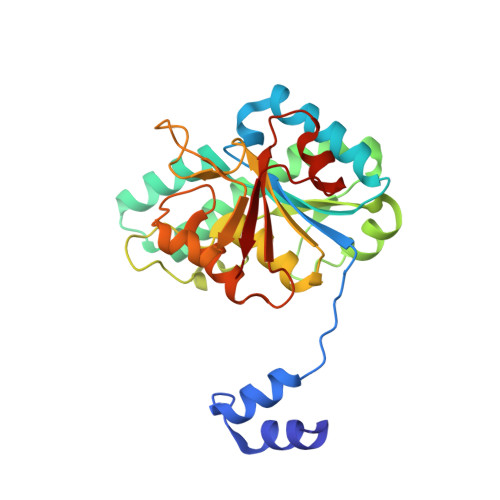
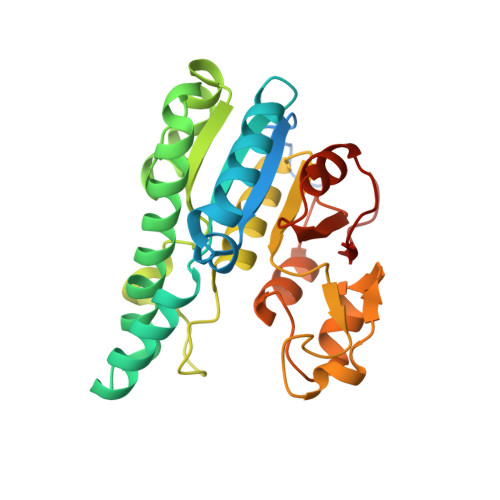
The molybdenum storage protein - A bionanolab for creating experimentally alterable polyoxomolybdate clusters.
Brunle, S., Poppe, J., Hail, R., Demmer, U., Ermler, U.(2018) J Inorg Biochem 189: 172-179
- PubMed: 30278367 Search on PubMed
- DOI: https://doi.org/10.1016/j.jinorgbio.2018.09.011
- Primary Citation Related Structures:
6GWB, 6H6W, 6H73, 6H74, 6H8B, 6H8H - Max-Planck-Institut für Biophysik, Max-von-Laue-Str. 3, D-60438 Frankfurt/Main, Germany.
Organizational Affiliation: