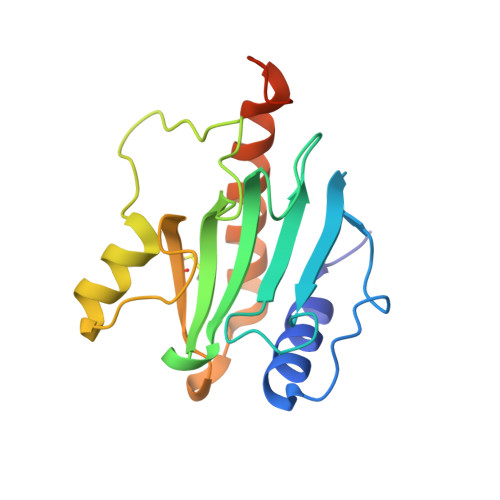
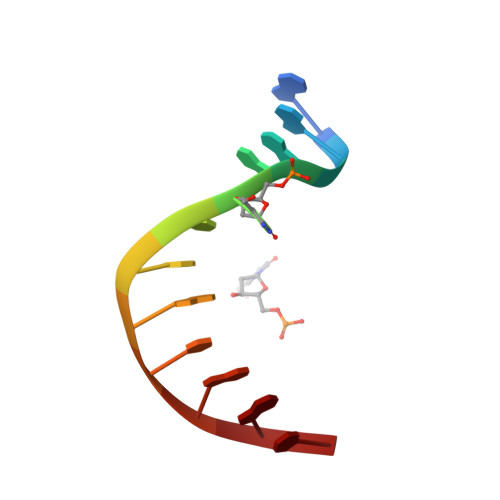
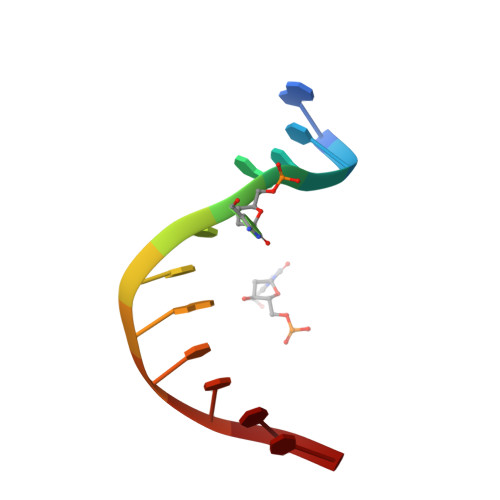
Recognition of modified cytosine variants by the DNA-binding domain of methyl-directed endonuclease McrBC.
Zagorskaite, E., Manakova, E., Sasnauskas, G.(2018) FEBS Lett 592: 3335-3345
- PubMed: 30194838 Search on PubMed
- DOI: https://doi.org/10.1002/1873-3468.13244
- Primary Citation Related Structures:
6GCD, 6GCE, 6GCF - PubMed Abstract:
Cytosine modifications expand the information content of genomic DNA in both eukaryotes and prokaryotes, providing means for epigenetic regulation and self versus nonself discrimination. For example, the methyl-directed restriction endonuclease, McrBC, recognizes and cuts invading bacteriophage DNA containing 5-methylcytosine (5mC), 5-hydroxymethylcytosine (5hmC), and N4-methylcytosine (4mC), leaving the unmodified host DNA intact. Here, we present cocrystal structures of McrB-N bound to DNA oligoduplexes containing 5hmC, 5-formylcytosine (5fC), and 4mC, and characterize the relative affinity of McrB-N to various cytosine variants. We find that McrB-N flips out modified bases into a protein pocket and binds cytosine derivatives in the order of descending affinity: 4mC > 5mC > 5hmC ≫ 5fC. We also show that pocket mutations alter the relative preference of McrB-N to 5mC, 5hmC, and 4mC.
- Institute of Biotechnology, Vilnius University, Vilnius, Lithuania.
Organizational Affiliation: