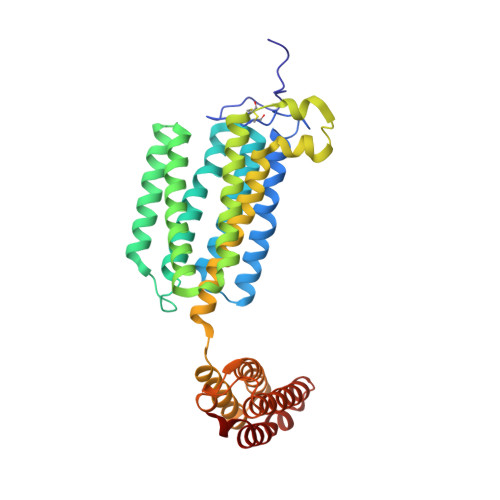
Structure of a human intramembrane ceramidase explains enzymatic dysfunction found in leukodystrophy.
Vasiliauskaite-Brooks, I., Healey, R.D., Rochaix, P., Saint-Paul, J., Sounier, R., Grison, C., Waltrich-Augusto, T., Fortier, M., Hoh, F., Saied, E.M., Arenz, C., Basu, S., Leyrat, C., Granier, S.(2018) Nat Commun 9: 5437-5437
- PubMed: 30575723 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-07864-w
- Primary Citation Related Structures:
6G7O - PubMed Abstract:
Alkaline ceramidases (ACERs) are a class of poorly understood transmembrane enzymes controlling the homeostasis of ceramides. They are implicated in human pathophysiology, including progressive leukodystrophy, colon cancer as well as acute myeloid leukemia. We report here the crystal structure of the human ACER type 3 (ACER3). Together with computational studies, the structure reveals that ACER3 is an intramembrane enzyme with a seven transmembrane domain architecture and a catalytic Zn 2+ binding site in its core, similar to adiponectin receptors. Interestingly, we uncover a Ca 2+ binding site physically and functionally connected to the Zn 2+ providing a structural explanation for the known regulatory role of Ca 2+ on ACER3 enzymatic activity and for the loss of function in E33G-ACER3 mutant found in leukodystrophic patients.
- IGF, University of Montpellier, CNRS, INSERM, Montpellier, 34094, France.
Organizational Affiliation: