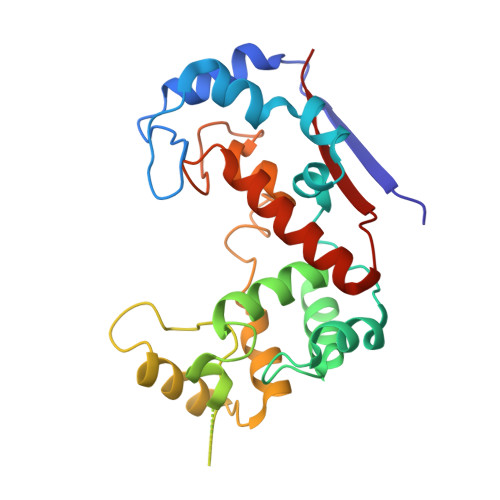
Domain sliding of two Staphylococcus aureus N-acetylglucosaminidases enables their substrate-binding prior to its catalysis.
Pintar, S., Borisek, J., Usenik, A., Perdih, A., Turk, D.(2020) Commun Biol 3: 178-178
- PubMed: 32313083 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-020-0911-7
- Primary Citation Related Structures:
6FXO, 6FXP - PubMed Abstract:
To achieve productive binding, enzymes and substrates must align their geometries to complement each other along an entire substrate binding site, which may require enzyme flexibility. In pursuit of novel drug targets for the human pathogen S. aureus, we studied peptidoglycan N-acetylglucosaminidases, whose structures are composed of two domains forming a V-shaped active site cleft. Combined insights from crystal structures supported by site-directed mutagenesis, modeling, and molecular dynamics enabled us to elucidate the substrate binding mechanism of SagB and AtlA-gl. This mechanism requires domain sliding from the open form observed in their crystal structures, leading to polysaccharide substrate binding in the closed form, which can enzymatically process the bound substrate. We suggest that these two hydrolases must exhibit unusual extents of flexibility to cleave the rigid structure of a bacterial cell wall.
- Department of Biochemistry, Molecular and Structural Biology, Jožef Stefan Institute, Jamova cesta 39, 1000, Ljubljana, Slovenia.
Organizational Affiliation: