Characterization of a potent and highly unusual minimally enhancing antibody directed against dengue virus.
Renner, M., Flanagan, A., Dejnirattisai, W., Puttikhunt, C., Kasinrerk, W., Supasa, P., Wongwiwat, W., Chawansuntati, K., Duangchinda, T., Cowper, A., Midgley, C.M., Malasit, P., Huiskonen, J.T., Mongkolsapaya, J., Screaton, G.R., Grimes, J.M.(2018) Nat Immunol 19: 1248-1256
- PubMed: 30323338 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41590-018-0227-7
- Primary Citation Related Structures:
6FLA, 6FLB, 6FLC - PubMed Abstract:
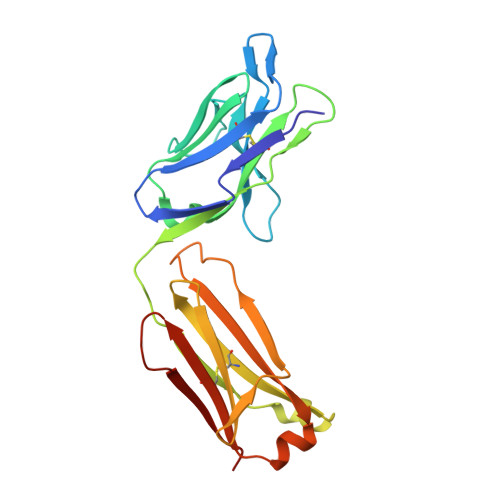
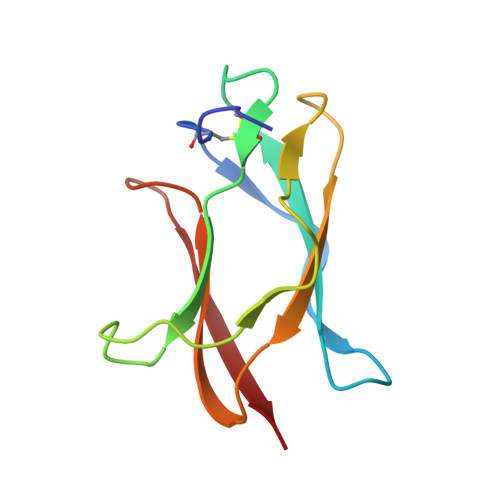
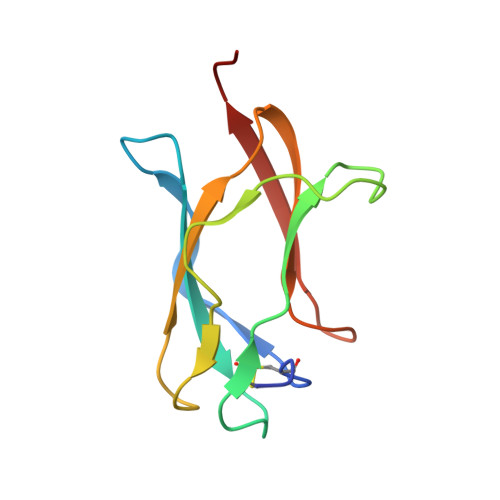
Dengue virus is a major pathogen, and severe infections can lead to life-threatening dengue hemorrhagic fever. Dengue virus exists as four serotypes, and dengue hemorrhagic fever is often associated with secondary heterologous infections. Antibody-dependent enhancement (ADE) may drive higher viral loads in these secondary infections and is purported to result from antibodies that recognize dengue virus but fail to fully neutralize it. Here we characterize two antibodies, 2C8 and 3H5, that bind to the envelope protein. Antibody 3H5 is highly unusual as it not only is potently neutralizing but also promotes little if any ADE, whereas antibody 2C8 has strong capacity to promote ADE. We show that 3H5 shows resilient binding in endosomal pH conditions and neutralizes at low occupancy. Immunocomplexes of 3H5 and dengue virus do not efficiently interact with Fcγ receptors, which we propose is due to the binding mode of 3H5 and constitutes the primary mechanism of how ADE is avoided.
- Division of Structural Biology, Wellcome Trust Centre for Human Genetics, University of Oxford, Oxford, UK.
Organizational Affiliation: