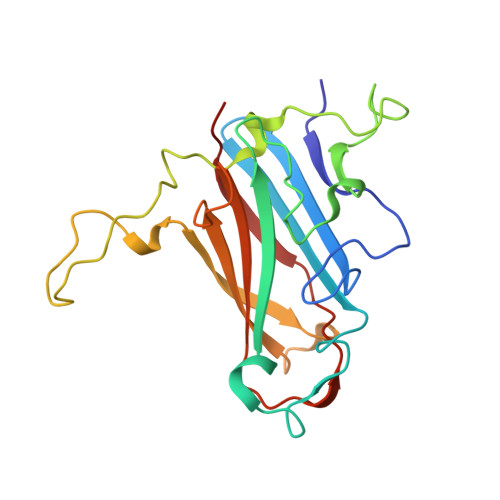
Diversity within the adenovirus fiber knob hypervariable loops influences primary receptor interactions.
Baker, A.T., Greenshields-Watson, A., Coughlan, L., Davies, J.A., Uusi-Kerttula, H., Cole, D.K., Rizkallah, P.J., Parker, A.L.(2019) Nat Commun 10: 741-741
- PubMed: 30765704 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-08599-y
- Primary Citation Related Structures:
6FJN, 6FJQ, 6HCN - PubMed Abstract:
Adenovirus based vectors are of increasing importance for wide ranging therapeutic applications. As vaccines, vectors derived from human adenovirus species D serotypes 26 and 48 (HAdV-D26/48) are demonstrating promising efficacy as protective platforms against infectious diseases. Significant clinical progress has been made, yet definitive studies underpinning mechanisms of entry, infection, and receptor usage are currently lacking. Here, we perform structural and biological analysis of the receptor binding fiber-knob protein of HAdV-D26/48, reporting crystal structures, and modelling putative interactions with two previously suggested attachment receptors, CD46 and Coxsackie and Adenovirus Receptor (CAR). We provide evidence of a low affinity interaction with CAR, with modelling suggesting affinity is attenuated through extended, semi-flexible loop structures, providing steric hindrance. Conversely, in silico and in vitro experiments are unable to provide evidence of interaction between HAdV-D26/48 fiber-knob with CD46, or with Desmoglein 2. Our findings provide insight into the cell-virus interactions of HAdV-D26/48, with important implications for the design and engineering of optimised Ad-based therapeutics.
- Division of Cancer and Genetics, School of Medicine, Cardiff University, Cardiff, CF14 4XN, UK.
Organizational Affiliation: