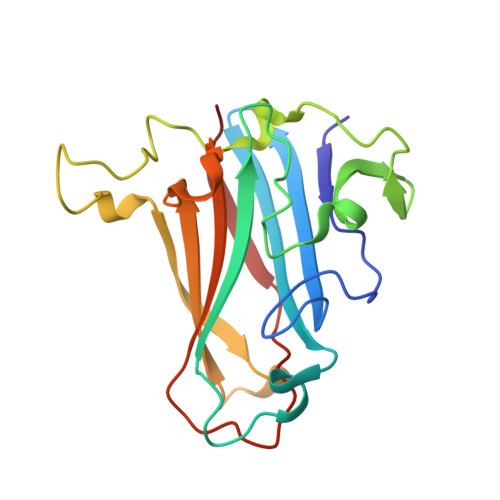
Human adenovirus type 26 uses sialic acid-bearing glycans as a primary cell entry receptor.
Baker, A.T., Mundy, R.M., Davies, J.A., Rizkallah, P.J., Parker, A.L.(2019) Sci Adv 5: eaax3567-eaax3567
- PubMed: 31517055 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.aax3567
- Primary Citation Related Structures:
6FJO, 6QU6, 6QU8 - PubMed Abstract:
Adenoviruses are clinically important agents. They cause respiratory distress, gastroenteritis, and epidemic keratoconjunctivitis. As non-enveloped, double-stranded DNA viruses, they are easily manipulated, making them popular vectors for therapeutic applications, including vaccines. Species D adenovirus type 26 (HAdV-D26) is both a cause of EKC and other diseases and a promising vaccine vector. HAdV-D26-derived vaccines are under investigation as protective platforms against HIV, Zika, and respiratory syncytial virus infections and are in phase 3 clinical trials for Ebola. We recently demonstrated that HAdV-D26 does not use CD46 or Desmoglein-2 as entry receptors, while the putative interaction with coxsackie and adenovirus receptor is low affinity and unlikely to represent the primary cell receptor. Here, we establish sialic acid as a primary entry receptor used by HAdV-D26. We demonstrate that removal of cell surface sialic acid inhibits HAdV-D26 infection, and provide a high-resolution crystal structure of HAdV-D26 fiber-knob in complex with sialic acid.
- Division of Cancer and Genetics, School of Medicine, Cardiff University, Heath Park, Cardiff CF14 4XN, UK.
Organizational Affiliation: