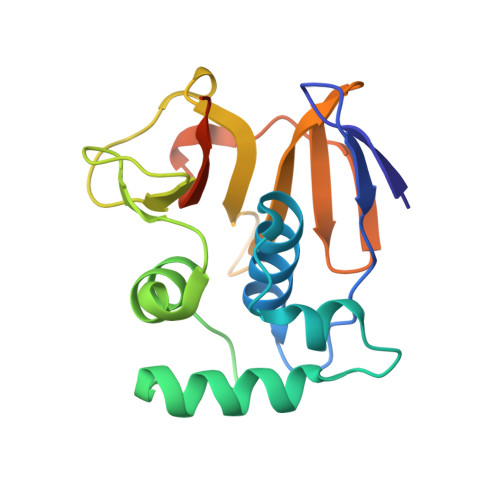
Irreversible inactivation of ISG15 by a viral leader protease enables alternative infection detection strategies.
Swatek, K.N., Aumayr, M., Pruneda, J.N., Visser, L.J., Berryman, S., Kueck, A.F., Geurink, P.P., Ovaa, H., van Kuppeveld, F.J.M., Tuthill, T.J., Skern, T., Komander, D.(2018) Proc Natl Acad Sci U S A 115: 2371-2376
- PubMed: 29463763 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1710617115
- Primary Citation Related Structures:
6FFA - PubMed Abstract:
In response to viral infection, cells mount a potent inflammatory response that relies on ISG15 and ubiquitin posttranslational modifications. Many viruses use deubiquitinases and deISGylases that reverse these modifications and antagonize host signaling processes. We here reveal that the leader protease, Lb pro , from foot-and-mouth disease virus (FMDV) targets ISG15 and to a lesser extent, ubiquitin in an unprecedented manner. Unlike canonical deISGylases that hydrolyze the isopeptide linkage after the C-terminal GlyGly motif, Lb pro cleaves the peptide bond preceding the GlyGly motif. Consequently, the GlyGly dipeptide remains attached to the substrate Lys, and cleaved ISG15 is rendered incompetent for reconjugation. A crystal structure of Lb pro bound to an engineered ISG15 suicide probe revealed the molecular basis for ISG15 proteolysis. Importantly, anti-GlyGly antibodies, developed for ubiquitin proteomics, are able to detect Lb pro cleavage products during viral infection. This opens avenues for infection detection of FMDV based on an immutable, host-derived epitope.
- Protein and Nucleic Acid Chemistry Division, Medical Research Council Laboratory of Molecular Biology, CB2 0QH Cambridge, United Kingdom.
Organizational Affiliation: