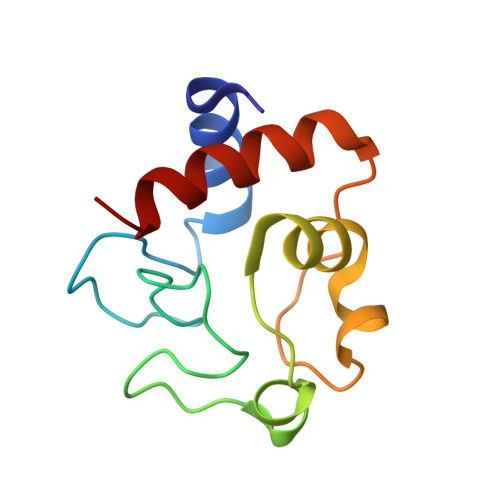
X-ray structure of bovine heart cytochrome c at high ionic strength.
Merlino, A.(2018) Biometals 31: 277-284
- PubMed: 29516298 Search on PubMed
- DOI: https://doi.org/10.1007/s10534-018-0090-x
- Primary Citation Related Structures:
6FF5 - PubMed Abstract:
Bovine heart cytochrome c (bCyt c) is an extensively studied hemoprotein of only 104 residues. Due to the existence of isoforms generated by non-enzymatic deaminidation, crystallization of bCyt c is difficult and involves extensive purification and the use of microseeding or the presence of an electric field. Taking advantage of the capacity of cytochrome c (cyt c) to bind anions on its protein surface, the commercially available bCyt c was crystallized without extra purifications, using ammonium sulfate as precipitant and nitrate ions as additives. The structure of the ferric bCyt c in a new crystal form is described and compared with that previously solved at low ionic strength and with those of human and horse cyt c. The overall structure of bCyt c is conserved, while the side chains of several residues that play a role in the interactions of cyt c with its partners have different rotamers in the two structures. The effect of the presence of nitrate ions on the structure of the protein is then evaluated and compared with that observed in the case of ferrous and ferric horse heart cyt c.
- Department of Chemical Sciences, University of Naples Federico II, Complesso Universitario di Monte Sant'Angelo, Via Cintia, 80126, Naples, Italy. antonello.merlino@unina.it.
Organizational Affiliation: