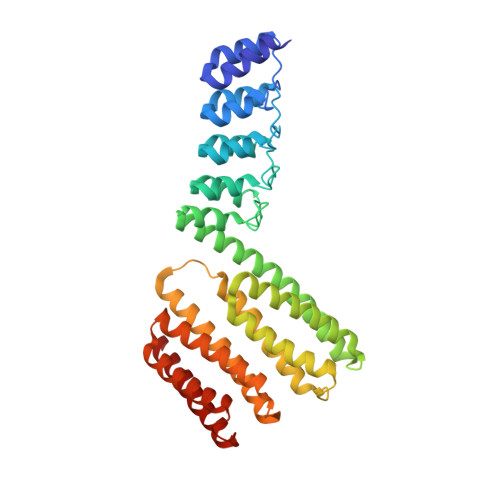
An Interface-Driven Design Strategy Yields a Novel, Corrugated Protein Architecture.
ElGamacy, M., Coles, M., Ernst, P., Zhu, H., Hartmann, M.D., Pluckthun, A., Lupas, A.N.(2018) ACS Synth Biol 7: 2226-2235
- PubMed: 30148951 Search on PubMed
- DOI: https://doi.org/10.1021/acssynbio.8b00224
- Primary Citation Related Structures:
6FES, 6FF6 - PubMed Abstract:
Designing proteins with novel folds remains a major challenge, as the biophysical properties of the target fold are not known a priori and no sequence profile exists to describe its features. Therefore, most computational design efforts so far have been directed toward creating proteins that recapitulate existing folds. Here we present a strategy centered upon the design of novel intramolecular interfaces that enables the construction of a target fold from a set of starting fragments. This strategy effectively reduces the amount of computational sampling necessary to achieve an optimal sequence, without compromising the level of topological control. The solenoid architecture has been a target of extensive protein design efforts, as it provides a highly modular platform of low topological complexity. However, none of the previous efforts have attempted to depart from the natural form, which is characterized by a uniformly handed superhelical architecture. Here we aimed to design a more complex platform, abolishing the superhelicity by introducing internally alternating handedness, resulting in a novel, corrugated architecture. We employed our interface-driven strategy, designing three proteins and confirming the design by solving the structure of two examples.
- Department of Protein Evolution , Max-Planck-Institute for Developmental Biology , 72076 Tübingen , Germany.
Organizational Affiliation: