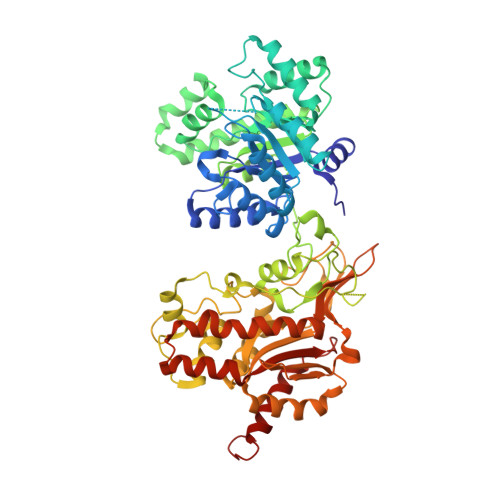
Structural basis for the regulation of human 5,10-methylenetetrahydrofolate reductase by phosphorylation and S-adenosylmethionine inhibition.
Froese, D.S., Kopec, J., Rembeza, E., Bezerra, G.A., Oberholzer, A.E., Suormala, T., Lutz, S., Chalk, R., Borkowska, O., Baumgartner, M.R., Yue, W.W.(2018) Nat Commun 9: 2261-2261
- PubMed: 29891918 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-04735-2
- Primary Citation Related Structures:
6FCX, 6FNU - PubMed Abstract:
The folate and methionine cycles are crucial for biosynthesis of lipids, nucleotides and proteins, and production of the methyl donor S-adenosylmethionine (SAM). 5,10-methylenetetrahydrofolate reductase (MTHFR) represents a key regulatory connection between these cycles, generating 5-methyltetrahydrofolate for initiation of the methionine cycle, and undergoing allosteric inhibition by its end product SAM. Our 2.5 Å resolution crystal structure of human MTHFR reveals a unique architecture, appending the well-conserved catalytic TIM-barrel to a eukaryote-only SAM-binding domain. The latter domain of novel fold provides the predominant interface for MTHFR homo-dimerization, positioning the N-terminal serine-rich phosphorylation region near the C-terminal SAM-binding domain. This explains how MTHFR phosphorylation, identified on 11 N-terminal residues (16 in total), increases sensitivity to SAM binding and inhibition. Finally, we demonstrate that the 25-amino-acid inter-domain linker enables conformational plasticity and propose it to be a key mediator of SAM regulation. Together, these results provide insight into the molecular regulation of MTHFR.
- Division of Metabolism and Children's Research Center, University Children's Hospital, CH-8032, Zürich, Switzerland. sean.froese@kispi.uzh.ch.
Organizational Affiliation: