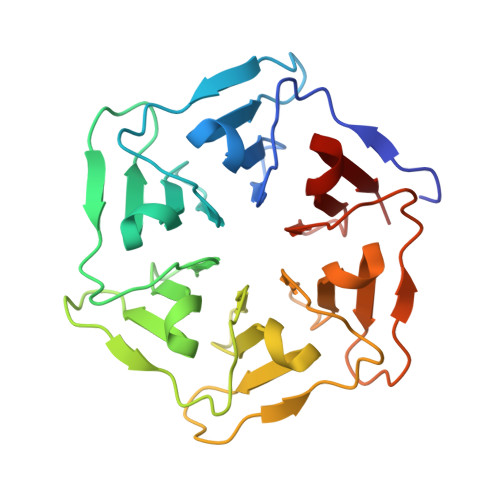
Design of tryptophan-containing mutants of the symmetrical Pizza protein for biophysical studies.
Noguchi, H., Mylemans, B., De Zitter, E., Van Meervelt, L., Tame, J.R.H., Voet, A.(2018) Biochem Biophys Res Commun 497: 1038-1042
- PubMed: 29481797 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2018.02.168
- Primary Citation Related Structures:
6F0Q, 6F0S, 6F0T - PubMed Abstract:
β-propeller proteins are highly symmetrical, being composed of a repeated motif with four anti-parallel β-sheets arranged around a central axis. Recently we designed the first completely symmetrical β-propeller protein, Pizza6, consisting of six identical tandem repeats. Pizza6 is expected to prove a useful building block for bionanotechnology, and also a tool to investigate the folding and evolution of β-propeller proteins. Folding studies are made difficult by the high stability and the lack of buried Trp residues to act as monitor fluorophores, so we have designed and characterized several Trp-containing Pizza6 derivatives. In total four proteins were designed, of which three could be purified and characterized. Crystal structures confirm these mutant proteins maintain the expected structure, and a clear redshift of Trp fluorescence emission could be observed upon denaturation. Among the derivative proteins, Pizza6-AYW appears to be the most suitable model protein for future folding/unfolding kinetics studies as it has a comparable stability as natural β-propeller proteins.
- Laboratory of Biomolecular Modelling and Design, Department of Chemistry, University of Leuven, Celestijnenlaan 200G-bus2403, Heverlee, Belgium.
Organizational Affiliation: