Design on a Rational Basis of High-Affinity Peptides Inhibiting the Histone Chaperone ASF1.
Bakail, M., Gaubert, A., Andreani, J., Moal, G., Pinna, G., Boyarchuk, E., Gaillard, M.C., Courbeyrette, R., Mann, C., Thuret, J.Y., Guichard, B., Murciano, B., Richet, N., Poitou, A., Frederic, C., Le Du, M.H., Agez, M., Roelants, C., Gurard-Levin, Z.A., Almouzni, G., Cherradi, N., Guerois, R., Ochsenbein, F.(2019) Cell Chem Biol 26: 1573-1585.e10
- PubMed: 31543461 Search on PubMed
- DOI: https://doi.org/10.1016/j.chembiol.2019.09.002
- Primary Citation Related Structures:
6F0F, 6F0G, 6F0H - PubMed Abstract:
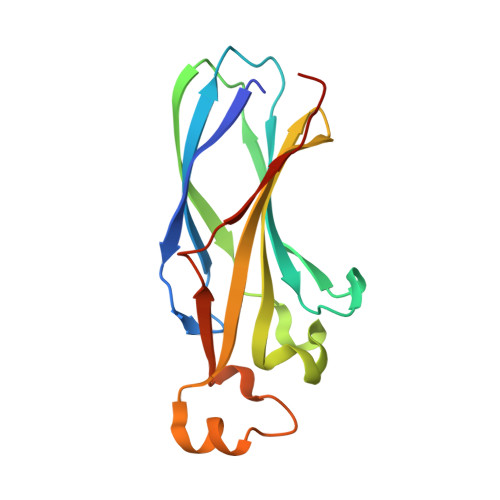
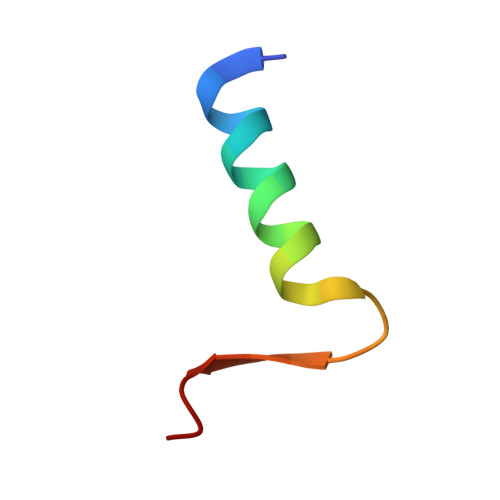
Anti-silencing function 1 (ASF1) is a conserved H3-H4 histone chaperone involved in histone dynamics during replication, transcription, and DNA repair. Overexpressed in proliferating tissues including many tumors, ASF1 has emerged as a promising therapeutic target. Here, we combine structural, computational, and biochemical approaches to design peptides that inhibit the ASF1-histone interaction. Starting from the structure of the human ASF1-histone complex, we developed a rational design strategy combining epitope tethering and optimization of interface contacts to identify a potent peptide inhibitor with a dissociation constant of 3 nM. When introduced into cultured cells, the inhibitors impair cell proliferation, perturb cell-cycle progression, and reduce cell migration and invasion in a manner commensurate with their affinity for ASF1. Finally, we find that direct injection of the most potent ASF1 peptide inhibitor in mouse allografts reduces tumor growth. Our results open new avenues to use ASF1 inhibitors as promising leads for cancer therapy.
- Institute Joliot, Commissariat à l'énergie Atomique (CEA), Direction de la Recherche Fondamentale (DRF), 91191 Gif-sur-Yvette, France; Institute for Integrative Biology of the Cell (I2BC), CEA, CNRS, Univ. Paris-Sud, Université Paris-Saclay, 91198 Gif-sur-Yvette Cedex, France.
Organizational Affiliation: