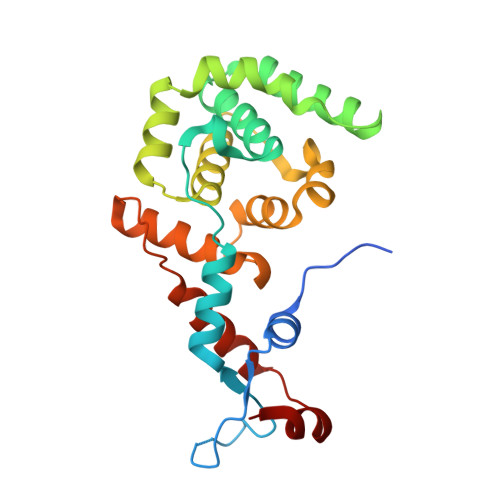
Legionella DotM structure reveals a role in effector recruiting to the Type 4B secretion system.
Meir, A., Chetrit, D., Liu, L., Roy, C.R., Waksman, G.(2018) Nat Commun 9: 507-507
- PubMed: 29410427 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-017-02578-x
- Primary Citation Related Structures:
6EXA, 6EXB, 6EXC, 6EXD, 6EXE - PubMed Abstract:
Legionella pneumophila, a causative agent of pneumonia, utilizes the Type 4B secretion (T4BS) system to translocate over 300 effectors into the host cell during infection. T4BS systems are encoded by a large gene cluster termed dot/icm, three components of which, DotL, DotM, and DotN, form the "coupling complex", which serves as a platform for recruitment of effector proteins. One class of effectors includes proteins containing Glu-rich/E-block sequences at their C terminus. However, the protein or region of the coupling complex mediating recruitment of such effectors is unknown. Here we present the crystal structure of DotM. This all alpha-helical structure exhibits patches of positively charged residues. We show that these regions form binding sites for acidic Glu-rich peptides and that mutants targeting these patches are defective in vivo in the translocation of acidic Glu-rich motif-containing effectors. We conclude that DotM forms the interacting surface for recruitment of acidic Glu-rich motif-containing Legionella effectors.
- Department of Biological Sciences, Institute of Structural and Molecular Biology, Birkbeck, Malet Street, London, WC1E 7HX, UK.
Organizational Affiliation: