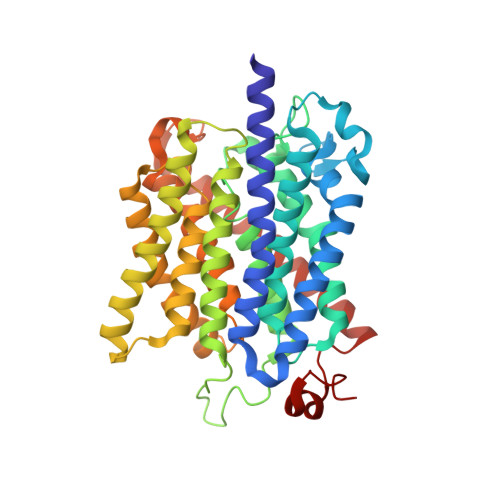
Signaling ammonium across membranes through an ammonium sensor histidine kinase.
Pfluger, T., Hernandez, C.F., Lewe, P., Frank, F., Mertens, H., Svergun, D., Baumstark, M.W., Lunin, V.Y., Jetten, M.S.M., Andrade, S.L.A.(2018) Nat Commun 9: 164-164
- PubMed: 29323112 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-017-02637-3
- Primary Citation Related Structures:
6EU6 - PubMed Abstract:
Sensing and uptake of external ammonium is essential for anaerobic ammonium-oxidizing (anammox) bacteria, and is typically the domain of the ubiquitous Amt/Rh ammonium transporters. Here, we report on the structure and function of an ammonium sensor/transducer from the anammox bacterium "Candidatus Kuenenia stuttgartiensis" that combines a membrane-integral ammonium transporter domain with a fused histidine kinase. It contains a high-affinity ammonium binding site not present in assimilatory Amt proteins. The levels of phosphorylated histidine in the kinase are coupled to the presence of ammonium, as conformational changes during signal recognition by the Amt module are transduced internally to modulate the kinase activity. The structural analysis of this ammonium sensor by X-ray crystallography and small-angle X-ray-scattering reveals a flexible, bipartite system that recruits a large uptake transporter as a sensory module and modulates its functionality to achieve a mechanistic coupling to a kinase domain in order to trigger downstream signaling events.
- Institute for Biochemistry, University of Freiburg, Albertstr. 21, Freiburg, 79104, Germany.
Organizational Affiliation: