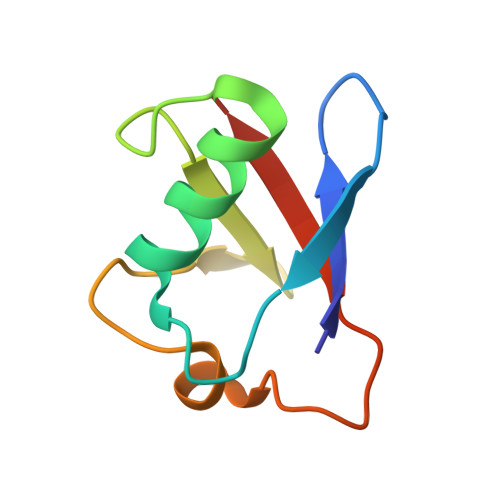
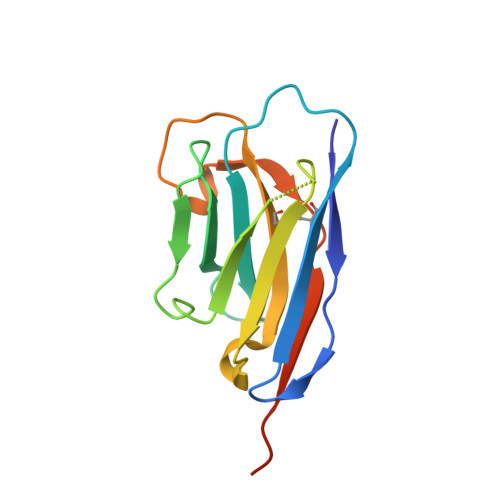
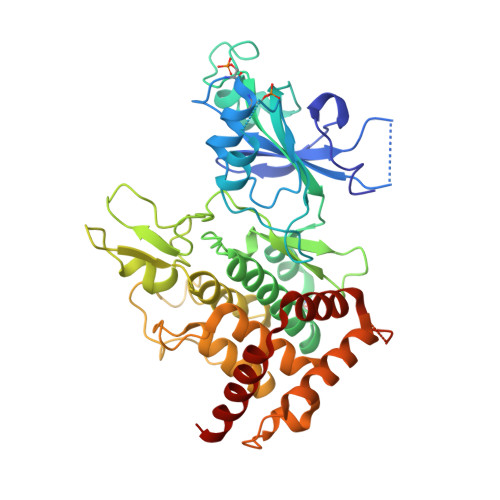
Structure of PINK1 in complex with its substrate ubiquitin.
Schubert, A.F., Gladkova, C., Pardon, E., Wagstaff, J.L., Freund, S.M.V., Steyaert, J., Maslen, S.L., Komander, D.(2017) Nature 552: 51-56
- PubMed: 29160309 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature24645
- Primary Citation Related Structures:
6EQI - PubMed Abstract:
Autosomal-recessive juvenile Parkinsonism (AR-JP) is caused by mutations in a number of PARK genes, in particular the genes encoding the E3 ubiquitin ligase Parkin (PARK2, also known as PRKN) and its upstream protein kinase PINK1 (also known as PARK6). PINK1 phosphorylates both ubiquitin and the ubiquitin-like domain of Parkin on structurally protected Ser65 residues, triggering mitophagy. Here we report a crystal structure of a nanobody-stabilized complex containing Pediculus humanus corporis (Ph)PINK1 bound to ubiquitin in the 'C-terminally retracted' (Ub-CR) conformation. The structure reveals many peculiarities of PINK1, including the architecture of the C-terminal region, and reveals how the N lobe of PINK1 binds ubiquitin via a unique insertion. The flexible Ser65 loop in the Ub-CR conformation contacts the activation segment, facilitating placement of Ser65 in a phosphate-accepting position. The structure also explains how autophosphorylation in the N lobe stabilizes structurally and functionally important insertions, and reveals the molecular basis of AR-JP-causing mutations, some of which disrupt ubiquitin binding.
- Medical Research Council Laboratory of Molecular Biology, Francis Crick Avenue, Cambridge CB2 0QH, UK.
Organizational Affiliation: