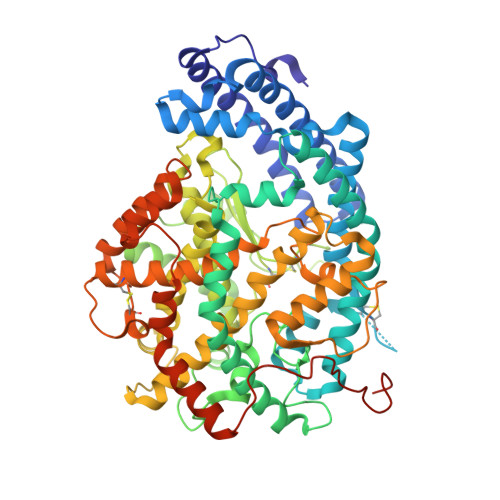
The Design and Development of a Potent and Selective Novel Diprolyl Derivative That Binds to the N-Domain of Angiotensin-I Converting Enzyme.
Fienberg, S., Cozier, G.E., Acharya, K.R., Chibale, K., Sturrock, E.D.(2018) J Med Chem 61: 344-359
- PubMed: 29206036 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.7b01478
- Primary Citation Related Structures:
6EN5, 6EN6 - PubMed Abstract:
Angiotensin-I converting enzyme (ACE) is a zinc metalloprotease consisting of two catalytic domains (N- and C-). Most clinical ACE inhibitor(s) (ACEi) have been shown to inhibit both domains nonselectively, resulting in adverse effects such as cough and angioedema. Selectively inhibiting the individual domains is likely to reduce these effects and potentially treat fibrosis in addition to hypertension. ACEi from the GVK Biosciences database were inspected for possible N-domain selective binding patterns. From this set, a diprolyl chemical series was modeled using docking simulations. The series was expanded based on key target interactions involving residues known to impart N-domain selectivity. In total, seven diprolyl compounds were synthesized and tested for N-domain selective ACE inhibition. One compound with an aspartic acid in the P 2 position (compound 16) displayed potent inhibition (K i = 11.45 nM) and was 84-fold more selective toward the N-domain. A high-resolution crystal structure of compound 16 in complex with the N-domain revealed the molecular basis for the observed selectivity.
- Department of Chemistry, University of Cape Town , Rondebosch 7701, South Africa.
Organizational Affiliation: