Structural Optimization of a Pyridinylimidazole Scaffold: Shifting the Selectivity from p38 alpha Mitogen-Activated Protein Kinase to c-Jun N-Terminal Kinase 3.
Ansideri, F., Macedo, J.T., Eitel, M., El-Gokha, A., Zinad, D.S., Scarpellini, C., Kudolo, M., Schollmeyer, D., Boeckler, F.M., Blaum, B.S., Laufer, S.A., Koch, P.(2018) ACS Omega 3: 7809-7831
- PubMed: 30087925 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsomega.8b00668
- Primary Citation Related Structures:
6EKD, 6EMH, 6EQ9 - PubMed Abstract:
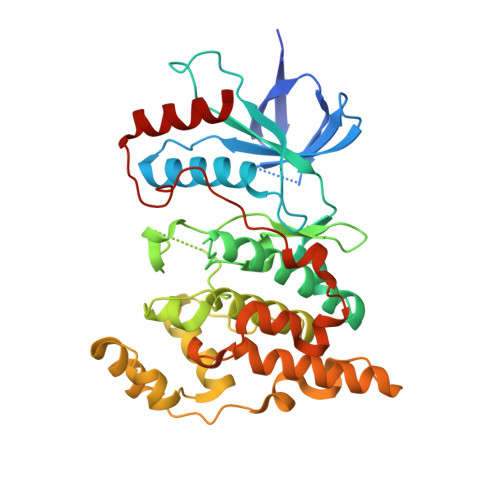
Starting from known p38α mitogen-activated protein kinase (MAPK) inhibitors, a series of inhibitors of the c-Jun N-terminal kinase (JNK) 3 was obtained. Altering the substitution pattern of the pyridinylimidazole scaffold proved to be effective in shifting the inhibitory activity from the original target p38α MAPK to the closely related JNK3. In particular, a significant improvement for JNK3 selectivity could be achieved by addressing the hydrophobic region I with a small methyl group. Furthermore, additional structural modifications permitted to explore structure-activity relationships. The most potent inhibitor 4-(4-methyl-2-(methylthio)-1 H -imidazol-5-yl)- N -(4-morpholinophenyl)pyridin-2-amine showed an IC 50 value for the JNK3 in the low triple digit nanomolar range and its binding mode was confirmed by X-ray crystallography.
- Department of Pharmaceutical and Medicinal Chemistry, Institute of Pharmaceutical Sciences, Eberhard Karls Universität Tübingen, Auf der Morgenstelle 8, 72076 Tübingen, Germany.
Organizational Affiliation: