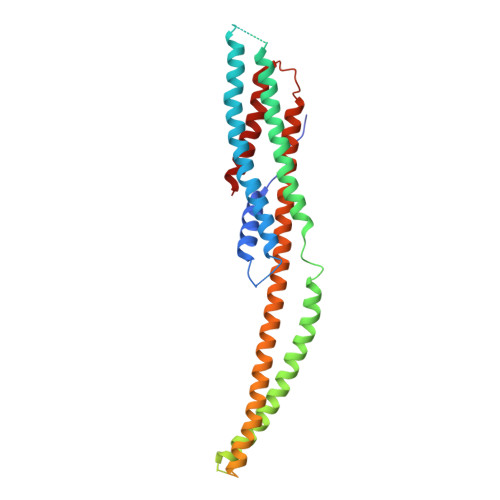
Structure and mechanism of the two-component alpha-helical pore-forming toxin YaxAB.
Brauning, B., Bertosin, E., Praetorius, F., Ihling, C., Schatt, A., Adler, A., Richter, K., Sinz, A., Dietz, H., Groll, M.(2018) Nat Commun 9: 1806-1806
- PubMed: 29728606 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-04139-2
- Primary Citation Related Structures:
6EK4, 6EK7, 6EK8, 6EL1 - PubMed Abstract:
Pore-forming toxins (PFT) are virulence factors that transform from soluble to membrane-bound states. The Yersinia YaxAB system represents a family of binary α-PFTs with orthologues in human, insect, and plant pathogens, with unknown structures. YaxAB was shown to be cytotoxic and likely involved in pathogenesis, though the molecular basis for its two-component lytic mechanism remains elusive. Here, we present crystal structures of YaxA and YaxB, together with a cryo-electron microscopy map of the YaxAB complex. Our structures reveal a pore predominantly composed of decamers of YaxA-YaxB heterodimers. Both subunits bear membrane-active moieties, but only YaxA is capable of binding to membranes by itself. YaxB can subsequently be recruited to membrane-associated YaxA and induced to present its lytic transmembrane helices. Pore formation can progress by further oligomerization of YaxA-YaxB dimers. Our results allow for a comparison between pore assemblies belonging to the wider ClyA-like family of α-PFTs, highlighting diverse pore architectures.
- Center for Integrated Protein Science Munich (CIPSM), Department of Chemistry, Chair of Biochemistry, Technische Universität München, Lichtenbergstrasse 4, 85747, Garching, Germany. bastian.braeuning@tum.de.
Organizational Affiliation: