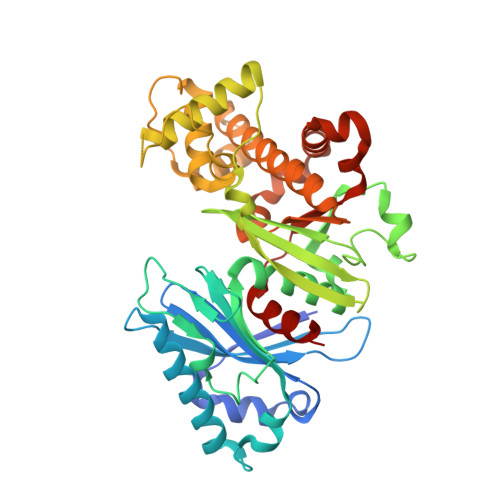
The crystal structure of glucokinase from Leishmania braziliensis.
Buechner, G.S., Millington, M.E., Perry, K., D'Antonio, E.L.(2019) Mol Biochem Parasitol 227: 47-52
- PubMed: 30571993 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molbiopara.2018.12.002
- Primary Citation Related Structures:
6EDI - PubMed Abstract:
Glucokinase from pathogenic protozoa of the genus Leishmania is a potential drug target for the chemotherapeutic treatment against leishmaniasis because this enzyme is located at a nodal point between two critically important metabolic pathways, glycolysis and the pentose phosphate pathway (PPP). L. braziliensis glucokinase (LbGlcK) was evaluated for its structural characterization and enzymatic performance. The enzyme catalyzes the phosphorylation of d-glucose with co-substrate ATP to yield the products G6P and ADP. LbGlcK had K M values determined as 6.61 ± 2.63 mM and 0.338 ± 0.080 mM for d-glucose and ATP, respectively. The 1.85 Å resolution X-ray crystal structure of the apo form of LbGlcK was determined and a homodimer was revealed where each subunit (both in open conformations) included the typical small and large domains. Structural comparisons were assessed in relationship to Homo sapiens hexokinase IV and Trypanosoma cruzi glucokinase. Comparisons revealed that all residues important for making hydrogen bonding interactions with d-glucose in the active site and catalysis were strictly conserved. LbGlcK was screened against four glucosamine analogue inhibitors and the stronger inhibitor of the series, HPOP-GlcN, had a K i value of 56.9 ± 16.6 μM that exhibited competitive inhibition. For the purpose of future structure-based drug design experimentation, L. braziliensis glucokinase was observed to be very similar to T. cruzi glucokinase even though there was a 44% protein sequence identity between the two enzymes.
- Department of Natural Sciences, University of South Carolina Beaufort, 1 University Boulevard, Bluffton, SC, 29909, USA.
Organizational Affiliation: