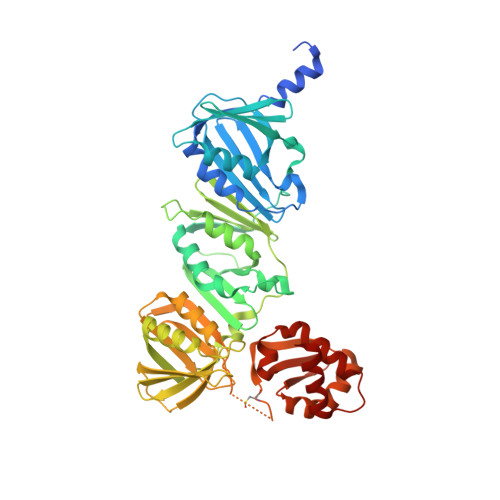
Binding of the regulatory domain of MutL to the sliding beta-clamp is species specific.
Almawi, A.W., Scotland, M.K., Randall, J.R., Liu, L., Martin, H.K., Sacre, L., Shen, Y., Pillon, M.C., Simmons, L.A., Sutton, M.D., Guarne, A.(2019) Nucleic Acids Res 47: 4831-4842
- PubMed: 30916336 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkz115
- Primary Citation Related Structures:
6E8D, 6E8E - PubMed Abstract:
The β-clamp is a protein hub central to DNA replication and fork management. Proteins interacting with the β-clamp harbor a conserved clamp-binding motif that is often found in extended regions. Therefore, clamp interactions have -almost exclusively- been studied using short peptides recapitulating the binding motif. This approach has revealed the molecular determinants that mediate the binding but cannot describe how proteins with clamp-binding motifs embedded in structured domains are recognized. The mismatch repair protein MutL has an internal clamp-binding motif, but its interaction with the β-clamp has different roles depending on the organism. In Bacillus subtilis, the interaction stimulates the endonuclease activity of MutL and it is critical for DNA mismatch repair. Conversely, disrupting the interaction between Escherichia coli MutL and the β-clamp only causes a mild mutator phenotype. Here, we determined the structures of the regulatory domains of E. coli and B. subtilis MutL bound to their respective β-clamps. The structures reveal different binding modes consistent with the binding to the β-clamp being a two-step process. Functional characterization indicates that, within the regulatory domain, only the clamp binding motif is required for the interaction between the two proteins. However, additional motifs beyond the regulatory domain may stabilize the interaction. We propose a model for the activation of the endonuclease activity of MutL in organisms lacking methyl-directed mismatch repair.
- Department of Biochemistry and Biomedical Sciences, McMaster University, Hamilton, ON, Canada.
Organizational Affiliation: