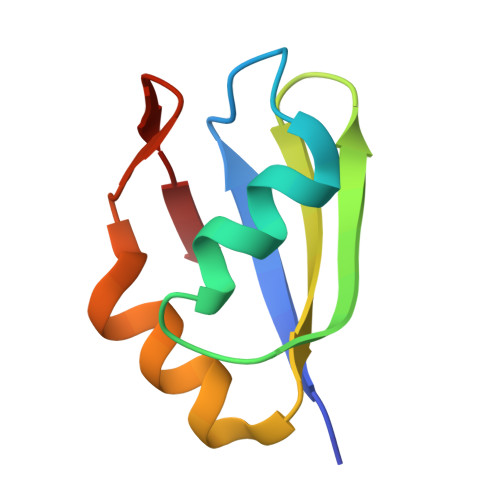
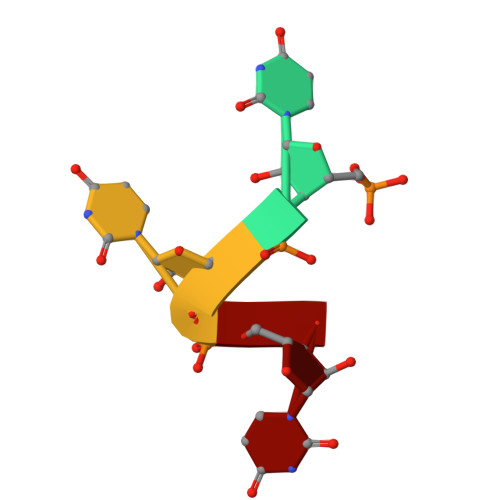
The RRM of the kRNA-editing protein TbRGG2 uses multiple surfaces to bind and remodel RNA.
Travis, B., Shaw, P.L.R., Liu, B., Ravindra, K., Iliff, H., Al-Hashimi, H.M., Schumacher, M.A.(2019) Nucleic Acids Res 47: 2130-2142
- PubMed: 30544166 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gky1259
- Primary Citation Related Structures:
6E4N, 6E4O, 6E4P - PubMed Abstract:
Kinetoplastid RNA (kRNA) editing takes place in the mitochondria of kinetoplastid protists and creates translatable mRNAs by uridine insertion/deletion. Extensively edited (pan-edited) transcripts contain quadruplex forming guanine stretches, which must be remodeled to promote uridine insertion/deletion. Here we show that the RRM domain of the essential kRNA-editing factor TbRGG2 binds poly(G) and poly(U) RNA and can unfold both. A region C-terminal to the RRM mediates TbRGG2 dimerization, enhancing RNA binding. A RRM-U4 RNA structure reveals a unique RNA-binding mechanism in which the two RRMs of the dimer employ aromatic residues outside the canonical RRM RNA-binding motifs to encase and wrench open the RNA, while backbone atoms specify the uridine bases. Notably, poly(G) RNA is bound via a different binding surface. Thus, these data indicate that TbRGG2 RRM can bind and remodel several RNA substrates suggesting how it might play multiple roles in the kRNA editing process.
- Department of Biochemistry, Duke University School of Medicine, Durham, NC 27710, USA.
Organizational Affiliation: