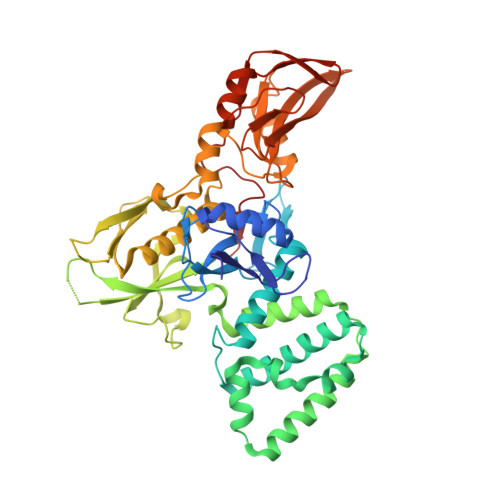
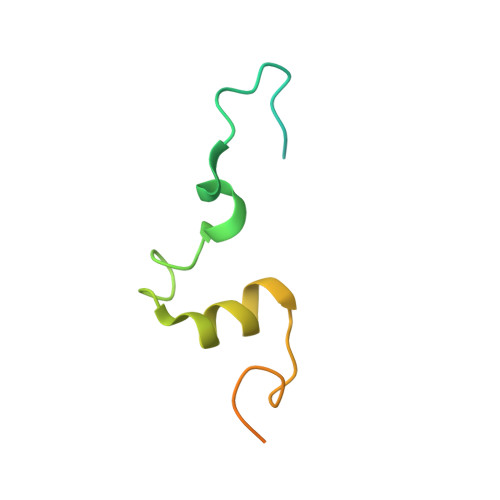
Receptor-mediated dimerization of JAK2 FERM domains is required for JAK2 activation.
Ferrao, R.D., Wallweber, H., Lupardus, P.J.(2018) Elife 7
- PubMed: 30044226 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.38089
- Primary Citation Related Structures:
6E2P, 6E2Q - PubMed Abstract:
Cytokines and interferons initiate intracellular signaling via receptor dimerization and activation of Janus kinases (JAKs). How JAKs structurally respond to changes in receptor conformation induced by ligand binding is not known. Here, we present two crystal structures of the human JAK2 FERM and SH2 domains bound to Leptin receptor (LEPR) and Erythropoietin receptor (EPOR), which identify a novel dimeric conformation for JAK2. This 2:2 JAK2/receptor dimer, observed in both structures, identifies a previously uncharacterized receptor interaction essential to dimer formation that is mediated by a membrane-proximal peptide motif called the 'switch' region. Mutation of the receptor switch region disrupts STAT phosphorylation but does not affect JAK2 binding, indicating that receptor-mediated formation of the JAK2 FERM dimer is required for kinase activation. These data uncover the structural and molecular basis for how a cytokine-bound active receptor dimer brings together two JAK2 molecules to stimulate JAK2 kinase activity.
- Department of Structural Biology, Genentech, Inc., South San Francisco, United States.
Organizational Affiliation: