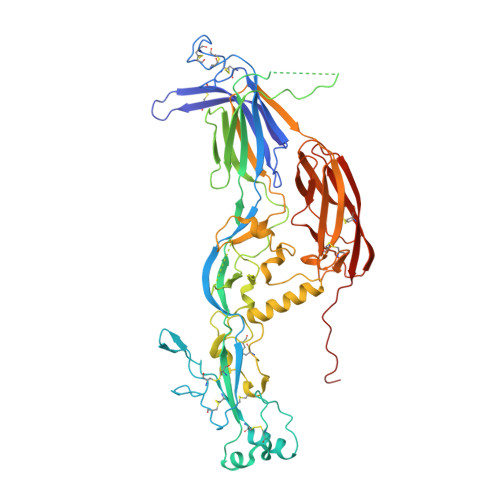
Species-Specific Functional Regions of the Green Alga Gamete Fusion Protein HAP2 Revealed by Structural Studies.
Baquero, E., Fedry, J., Legrand, P., Krey, T., Rey, F.A.(2019) Structure 27: 113
- PubMed: 30416037 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2018.09.014
- Primary Citation Related Structures:
6E18 - PubMed Abstract:
The cellular fusion protein HAP2, which is structurally homologous to viral class II fusion proteins, drives gamete fusion across several eukaryotic kingdoms. Gamete fusion is a highly controlled process in eukaryotes, and is allowed only between same species gametes. In spite of a conserved architecture, HAP2 displays several species-specific functional regions that were not resolved in the available X-ray structure of the green alga Chlamydomonas reinhardtii HAP2 ectodomain. Here we present an X-ray structure resolving these regions, showing a target membrane interaction surface made by three amphipathic helices in a horseshoe-shaped arrangement. HAP2 from green algae also features additional species-specific motifs inserted in regions that in viral class II proteins are critical for the fusogenic conformational change. Such insertions include a cystine ladder-like module evocative of EGF-like motifs responsible for extracellular protein-protein interactions in animals, and a mucin-like region. These features suggest potential HAP2 interaction sites involved in gamete fusion control.
- Institut Pasteur, Unité de Virologie Structurale, 25-28 Rue du Docteur Roux, 75724 Paris Cedex 15, France; CNRS UMR 3569, 25-28 Rue du Docteur Roux, 75724 Paris Cedex 15, France.
Organizational Affiliation: