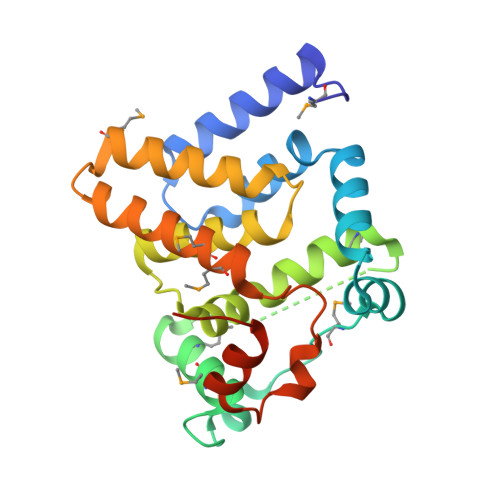
Crystal Structure of the Alr8543 protein in complex with Oleic Acid and magnesium ion from Nostoc sp. PCC 7120, Northeast Structural Genomics Consortium Target NsR141
Forouhar, F., Neely, H., Seetharaman, J., Xiao, R., Ciccosanti, C., Patel, D., Belote, R.L., Everett, J.K., Nair, R., Acton, T.B., Rost, B., Montelione, G.T., Tong, L., Hunt, J.F., Northeast Structural Genomics Consortium (NESG)To be published.