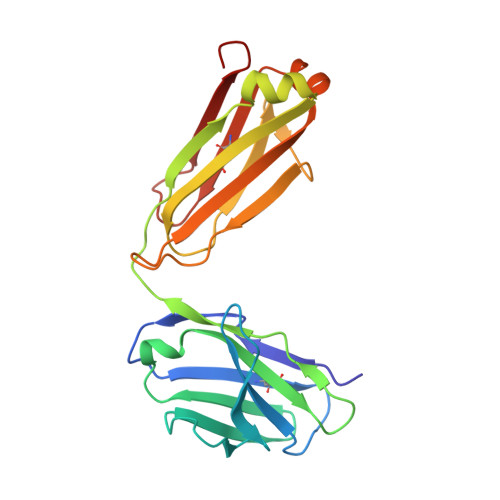
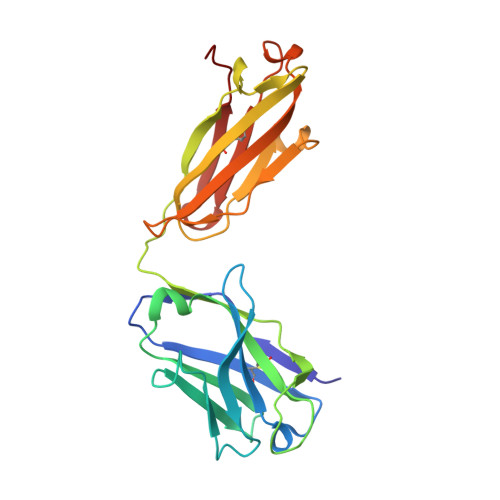
Structural Basis of Pan-Ebolavirus Neutralization by an Antibody Targeting the Glycoprotein Fusion Loop.
Murin, C.D., Bruhn, J.F., Bornholdt, Z.A., Copps, J., Stanfield, R., Ward, A.B.(2018) Cell Rep 24: 2723-2732.e4
- PubMed: 30184505 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2018.08.009
- Primary Citation Related Structures:
6DZL, 6DZM, 6DZN - PubMed Abstract:
Monoclonal antibodies (mAbs) with pan-ebolavirus cross-reactivity are highly desirable, but development of such mAbs is limited by a lack of a molecular understanding of cross-reactive epitopes. The antibody ADI-15878 was previously identified from a human survivor of Ebola virus Makona variant (EBOV/Mak) infection. This mAb demonstrated potent neutralizing activity against all known ebolaviruses and provided protection in rodent and ferret models against three ebolavirus species. Here, we describe the unliganded crystal structure of ADI-15878 as well as the cryo-EM structures of ADI-15878 in complex with the EBOV/Mak and Bundibugyo virus (BDBV) glycoproteins (GPs). ADI-15878 binds through an induced-fit mechanism by targeting highly conserved residues in the internal fusion loop (IFL), bridging across GP protomers via the heptad repeat 1 (HR1) region. Our structures provide a more complete description of the ebolavirus immunogenic landscape, as well as a molecular basis for how rare but potent antibodies target conserved filoviral fusion machinery.
- Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: