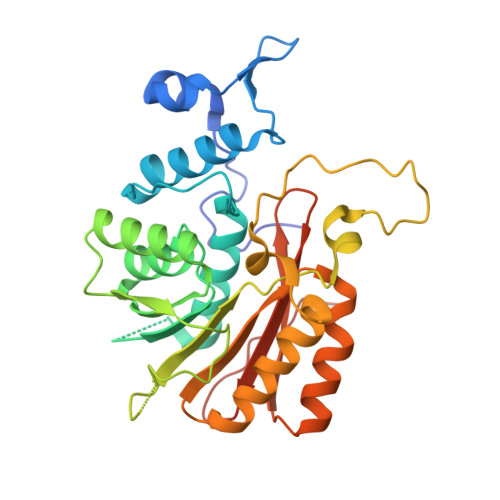
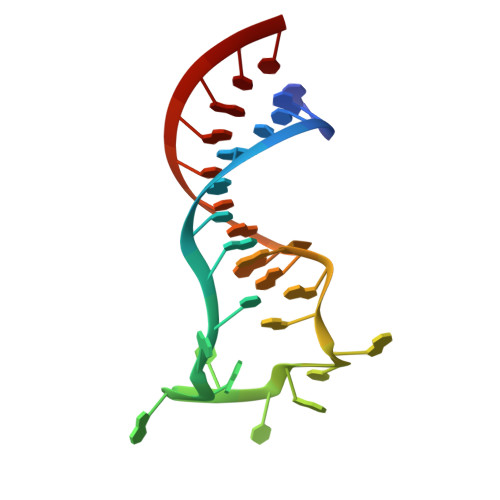
Structural Basis for Regulation of METTL16, an S-Adenosylmethionine Homeostasis Factor.
Doxtader, K.A., Wang, P., Scarborough, A.M., Seo, D., Conrad, N.K., Nam, Y.(2018) Mol Cell 71: 1001-1011.e4
- PubMed: 30197297 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2018.07.025
- Primary Citation Related Structures:
6DU4, 6DU5 - PubMed Abstract:
S-adenosylmethionine (SAM) is an essential metabolite that acts as a cofactor for most methylation events in the cell. The N 6 -methyladenosine (m 6 A) methyltransferase METTL16 controls SAM homeostasis by regulating the abundance of SAM synthetase MAT2A mRNA in response to changing intracellular SAM levels. Here we present crystal structures of METTL16 in complex with MAT2A RNA hairpins to uncover critical molecular mechanisms underlying the regulated activity of METTL16. The METTL16-RNA complex structures reveal atomic details of RNA substrates that drive productive methylation by METTL16. In addition, we identify a polypeptide loop in METTL16 near the SAM binding site with an autoregulatory role. We show that mutations that enhance or repress METTL16 activity in vitro correlate with changes in MAT2A mRNA levels in cells. Thus, we demonstrate the structural basis for the specific activity of METTL16 and further suggest the molecular mechanisms by which METTL16 efficiency is tuned to regulate SAM homeostasis.
- Cecil H. and Ida Green Center for Reproductive Biology Sciences and Division of Basic Reproductive Biology Research, Department of Obstetrics and Gynecology, University of Texas Southwestern Medical Center, Dallas, TX 75390, USA; Department of Biophysics, University of Texas Southwestern Medical Center, Dallas, TX 75390, USA.
Organizational Affiliation: