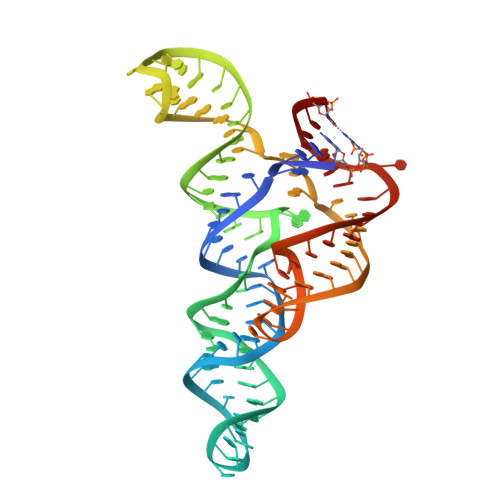
ykkC riboswitches employ an add-on helix to adjust specificity for polyanionic ligands.
Peselis, A., Serganov, A.(2018) Nat Chem Biol 14: 887-894
- PubMed: 30120360 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-018-0114-4
- Primary Citation Related Structures:
6DLQ, 6DLR, 6DLS, 6DLT, 6DMC, 6DMD, 6DME, 6DNR - PubMed Abstract:
The ykkC family of bacterial riboswitches combines several widespread classes that have similar secondary structures and consensus motifs but control different genes in response to different cellular metabolites. Here we report the crystal structures of two distinct ykkC riboswitches specifically bound to their cognate ligand ppGpp, a second messenger involved in stress response, or PRPP, a precursor in purine biosynthesis. Both RNAs adopt similar structures and contain a conserved core previously observed in the guanidine-specific ykkC riboswitch. However, ppGpp and PRPP riboswitches uniquely employ an additional helical element that joins the ends of the ligand-sensing domains and creates a tunnel for direct and Mg 2+ -mediated binding of ligands. Mutational and footprinting experiments highlight the importance of conserved nucleotides forming the tunnel and long-distance contacts for ligand binding and genetic response. Our work provides new insights into the specificity of riboswitches and gives a unique opportunity for future studies of RNA evolution.
- Department of Biochemistry and Molecular Pharmacology, New York University School of Medicine, New York, NY, USA.
Organizational Affiliation: