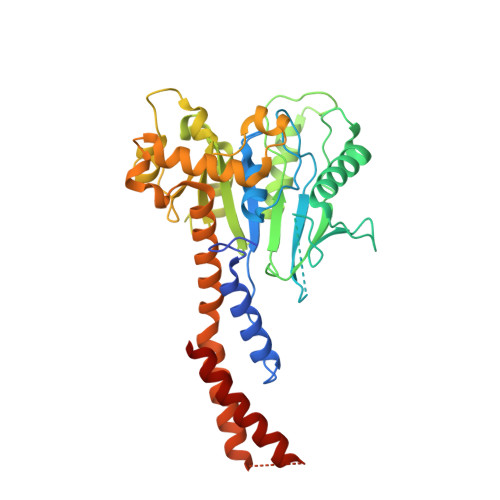
Structures of the fungal dynamin-related protein Vps1 reveal a unique, open helical architecture.
Varlakhanova, N.V., Alvarez, F.J.D., Brady, T.M., Tornabene, B.A., Hosford, C.J., Chappie, J.S., Zhang, P., Ford, M.G.J.(2018) J Cell Biol 217: 3608-3624
- PubMed: 30087125 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1083/jcb.201712021
- Primary Citation Related Structures:
6DEF, 6DI7, 6DJQ - PubMed Abstract:
Dynamin-related proteins (DRPs) are large multidomain GTPases required for diverse membrane-remodeling events. DRPs self-assemble into helical structures, but how these structures are tailored to their cellular targets remains unclear. We demonstrate that the fungal DRP Vps1 primarily localizes to and functions at the endosomal compartment. We present crystal structures of a Vps1 GTPase-bundle signaling element (BSE) fusion in different nucleotide states to capture GTP hydrolysis intermediates and concomitant conformational changes. Using cryoEM, we determined the structure of full-length GMPPCP-bound Vps1. The Vps1 helix is more open and flexible than that of dynamin. This is due to further opening of the BSEs away from the GTPase domains. A novel interface between adjacent GTPase domains forms in Vps1 instead of the contacts between the BSE and adjacent stalks and GTPase domains as seen in dynamin. Disruption of this interface abolishes Vps1 function in vivo. Hence, Vps1 exhibits a unique helical architecture, highlighting structural flexibilities of DRP self-assembly.
- Department of Cell Biology, University of Pittsburgh School of Medicine, Pittsburgh, PA.
Organizational Affiliation: