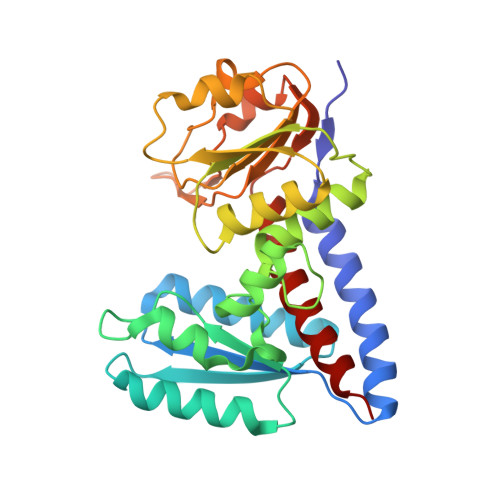
Crystal Structure of Bifunctional Enzyme FolD-Methylenetetrahydrofolate Dehydrogenase/Cyclohydrolase in the Complex with Methotrexate from Campylobacter jejuni
Kim, Y., Makowska-Grzyska, M., Maltseva, N., Grimshaw, S., Joachimiak, A., Center for Structural Genomics of Infectious Diseases (CSGID)To be published.