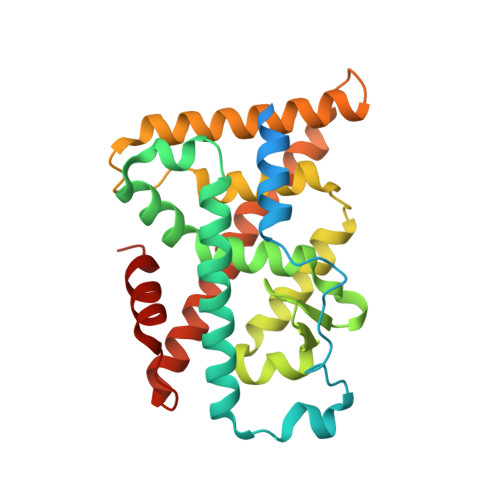
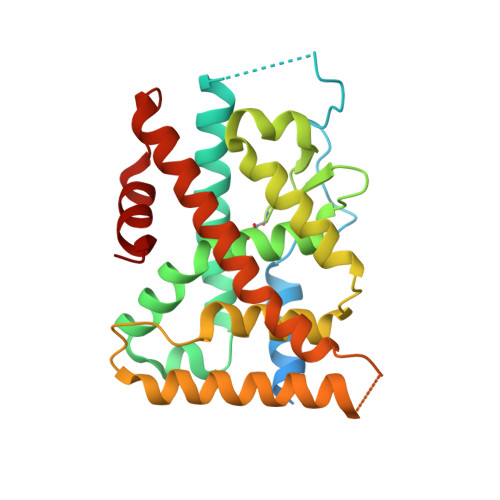
Covalent Modification and Regulation of the Nuclear Receptor Nurr1 by a Dopamine Metabolite.
Bruning, J.M., Wang, Y., Oltrabella, F., Tian, B., Kholodar, S.A., Liu, H., Bhattacharya, P., Guo, S., Holton, J.M., Fletterick, R.J., Jacobson, M.P., England, P.M.(2019) Cell Chem Biol 26: 674-685.e6
- PubMed: 30853418 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.chembiol.2019.02.002
- Primary Citation Related Structures:
6DDA - PubMed Abstract:
Nurr1, a nuclear receptor essential for the development, maintenance, and survival of midbrain dopaminergic neurons, is a potential therapeutic target for Parkinson's disease, a neurological disorder characterized by the degeneration of these same neurons. Efforts to identify Nurr1 agonists have been hampered by the recognition that it lacks several classic regulatory elements of nuclear receptor function, including the canonical ligand-binding pocket. Here we report that the dopamine metabolite 5,6-dihydroxyindole (DHI) binds directly to and modulates the activity of Nurr1. Using biophysical assays and X-ray crystallography, we show that DHI binds to the ligand-binding domain within a non-canonical pocket, forming a covalent adduct with Cys566. In cultured cells and zebrafish, DHI stimulates Nurr1 activity, including the transcription of target genes underlying dopamine homeostasis. These findings suggest avenues for developing synthetic Nurr1 ligands to ameliorate the symptoms and progression of Parkinson's disease.
- Pharmaceutical Sciences and Pharmacogenomics Graduate Program, University of California San Francisco, San Francisco, CA 94158, USA.
Organizational Affiliation: