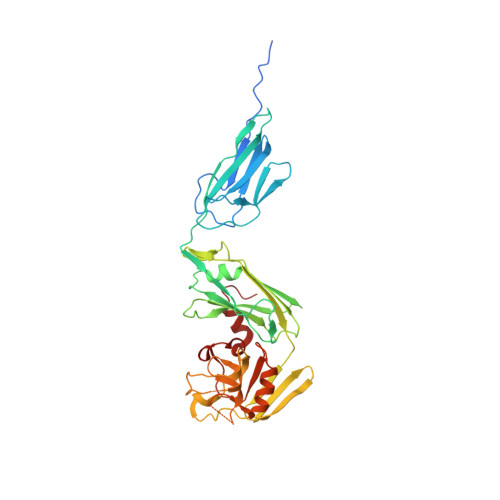
Structural Basis for the Interaction and Processing of beta-Lactam Antibiotics by l,d-Transpeptidase 3 (LdtMt3) from Mycobacterium tuberculosis.
Libreros-Zuniga, G.A., Dos Santos Silva, C., Salgado Ferreira, R., Dias, M.V.B.(2019) ACS Infect Dis 5: 260-271
- PubMed: 30556998 Search on PubMed
- DOI: https://doi.org/10.1021/acsinfecdis.8b00244
- Primary Citation Related Structures:
6D4K, 6D51, 6D5A - PubMed Abstract:
Targeting Mycobacterium tuberculosis peptidoglycans with β-lactam antibiotics represents a strategy to address increasing resistance to antitubercular drugs. β-Lactams inhibit peptidoglycan synthases such as l,d-transpeptidases, a group of carbapenem-sensitive enzymes that stabilize peptidoglycans through 3 → 3 cross-links. M. tuberculosis encodes five l,d-transpeptidases (Ldt Mt1-5 ), of which Ldt Mt3 is one of the less understood. Herein, we structurally characterized the apo and faropenem-acylated forms of Ldt Mt3 at 1.3 and 1.8 Å resolution, respectively. These structures revealed a fold and catalytic diad similar to those of other Ldts Mt enzymes, supporting its involvement in transpeptidation reactions despite divergences in active site size and charges. The Ldt Mt3 -faropenem structure indicated that faropenem is degraded after Cys-246 acylation, and possibly only a β-OH-butyrate or an acetyl group (C 2 H 3 O) covalently attached to the enzyme remains, an observation that strongly supports the notion that Ldt Mt3 is inactivated by β-lactams. Docking simulations with intact β-lactams predicted key Ldt Mt3 residues that interact with these antibiotics. We also characterized the heat of acylation involved in the binding and reaction of Ldt Mt3 for ten β-lactams belonging to four different classes, and imipenem had the highest inactivation constant. This work provides key insights into the structure, binding mechanisms, and degradation of β-lactams by Ldt Mt3 , which may be useful for the development of additional β-lactams with potential antitubercular activity.
- Departamento de Microbiologia, Instituto de Ciências Biomédicas , Universidade de São Paulo , Avenida Prof. Lineu Prestes , 1374 São Paulo , Brazil.
Organizational Affiliation: