A molecular switch in mouse CD1d modulates natural killer T cell activation by alpha-galactosylsphingamides.
Wang, J., Guillaume, J., Janssens, J., Remesh, S.G., Ying, G., Bitra, A., Van Calenbergh, S., Zajonc, D.M.(2019) J Biological Chem 294: 14345-14356
- PubMed: 31391251 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA119.009963
- Primary Citation Related Structures:
6C5M, 6C69, 6C6A, 6C6C, 6C6E, 6C6H, 6C6J, 6CW6, 6CW9, 6CWB, 6CX5, 6CX7, 6CX9, 6CXA, 6CXE, 6CXF, 6OJP - PubMed Abstract:
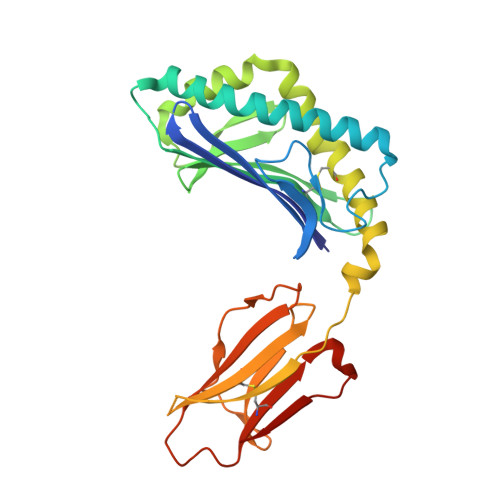
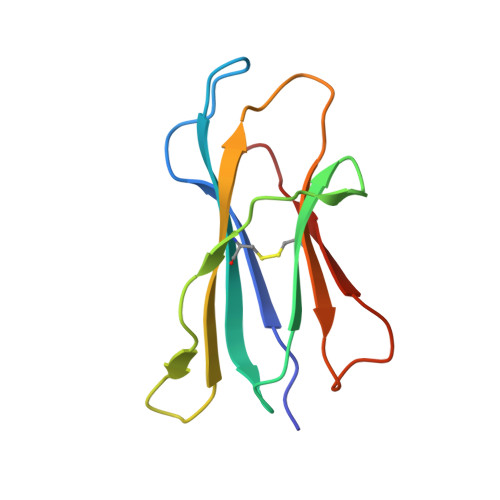
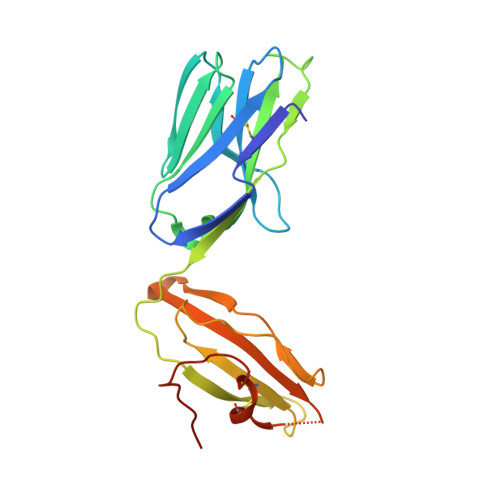
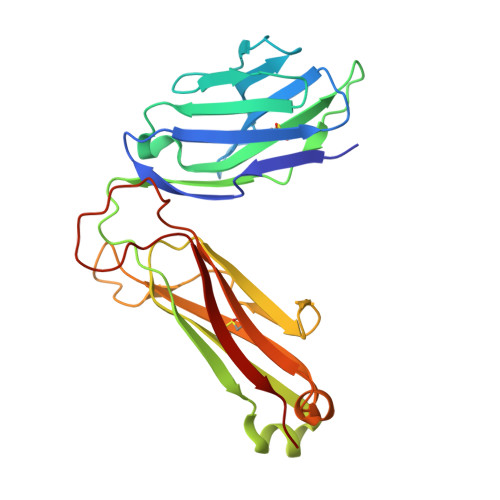
Type I natural killer T (NKT) cells are a population of innate like T lymphocytes that rapidly respond to α-GalCer presented by CD1d via the production of both pro- and anti-inflammatory cytokines. While developing novel α-GalCer analogs that were meant to be utilized as potential adjuvants because of their production of pro-inflammatory cytokines (Th1 skewers), we generated α-galactosylsphingamides (αGSA). Surprisingly, αGSAs are not potent antigens in vivo despite their strong T-cell receptor (TCR)-binding affinities. Here, using surface plasmon resonance (SPR), antigen presentation assays, and X-ray crystallography (yielding crystal structures of 19 different binary (CD1d-glycolipid) or ternary (CD1d-glycolipid-TCR) complexes at resolutions between 1.67 and 2.85 Å), we characterized the biochemical and structural details of αGSA recognition by murine NKT cells. We identified a molecular switch within murine (m)CD1d that modulates NKT cell activation by αGSAs. We found that the molecular switch involves a hydrogen bond interaction between Tyr-73 of mCD1d and the amide group oxygen of αGSAs. We further established that the length of the acyl chain controls the positioning of the amide group with respect to the molecular switch and works synergistically with Tyr-73 to control NKT cell activity. In conclusion, our findings reveal important mechanistic insights into the presentation and recognition of glycolipids with polar moieties in an otherwise apolar milieu. These observations may inform the development αGSAs as specific NKT cell antagonists to modulate immune responses.
- Division of Immune Regulation, La Jolla Institute for Immunology (LJI), La Jolla, California 92037.
Organizational Affiliation: