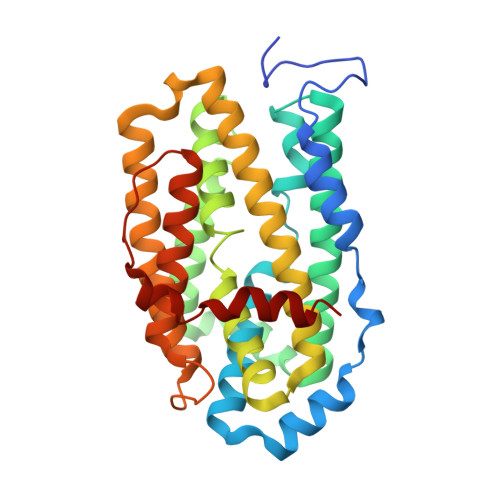
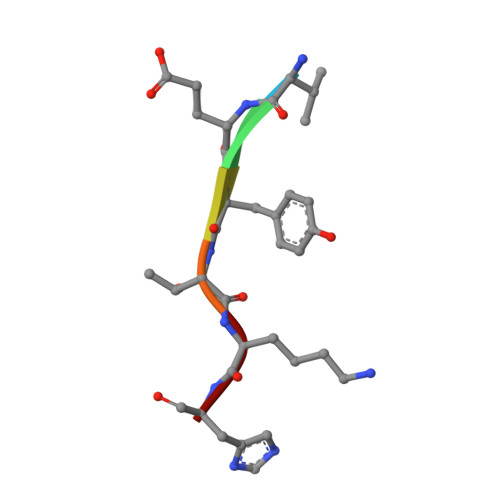
Structural Basis for Superoxide Activation of Flavobacterium johnsoniae Class I Ribonucleotide Reductase and for Radical Initiation by Its Dimanganese Cofactor.
Rose, H.R., Ghosh, M.K., Maggiolo, A.O., Pollock, C.J., Blaesi, E.J., Hajj, V., Wei, Y., Rajakovich, L.J., Chang, W.C., Han, Y., Hajj, M., Krebs, C., Silakov, A., Pandelia, M.E., Bollinger, J.M., Boal, A.K.(2018) Biochemistry 57: 2679-2693
- PubMed: 29609464 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.8b00247
- Primary Citation Related Structures:
6CWO, 6CWP, 6CWQ - PubMed Abstract:
A ribonucleotide reductase (RNR) from Flavobacterium johnsoniae ( Fj) differs fundamentally from known (subclass a-c) class I RNRs, warranting its assignment to a new subclass, Id. Its β subunit shares with Ib counterparts the requirements for manganese(II) and superoxide (O 2 - ) for activation, but it does not require the O 2 - -supplying flavoprotein (NrdI) needed in Ib systems, instead scavenging the oxidant from solution. Although Fj β has tyrosine at the appropriate sequence position (Tyr 104), this residue is not oxidized to a radical upon activation, as occurs in the Ia/b proteins. Rather, Fj β directly deploys an oxidized dimanganese cofactor for radical initiation. In treatment with one-electron reductants, the cofactor can undergo cooperative three-electron reduction to the II/II state, in contrast to the quantitative univalent reduction to inactive "met" (III/III) forms seen with I(a-c) βs. This tendency makes Fj β unusually robust, as the II/II form can readily be reactivated. The structure of the protein rationalizes its distinctive traits. A distortion in a core helix of the ferritin-like architecture renders the active site unusually open, introduces a cavity near the cofactor, and positions a subclass-d-specific Lys residue to shepherd O 2 - to the Mn 2 II/II cluster. Relative to the positions of the radical tyrosines in the Ia/b proteins, the unreactive Tyr 104 of Fj β is held away from the cofactor by a hydrogen bond with a subclass-d-specific Thr residue. Structural comparisons, considered with its uniquely simple mode of activation, suggest that the Id protein might most closely resemble the primordial RNR-β.
- Department of Chemistry , Massachusetts Institute of Technology , Cambridge , Massachusetts 02139 , United States.
Organizational Affiliation: