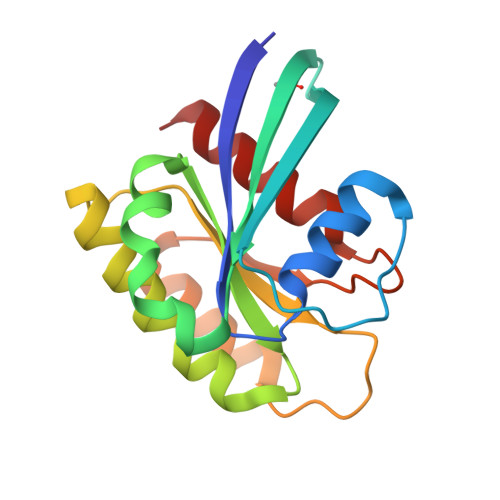
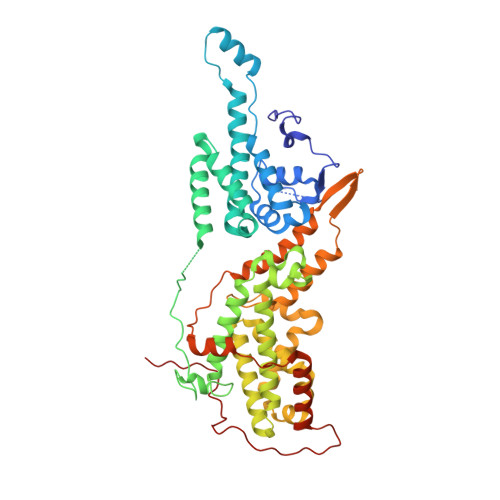
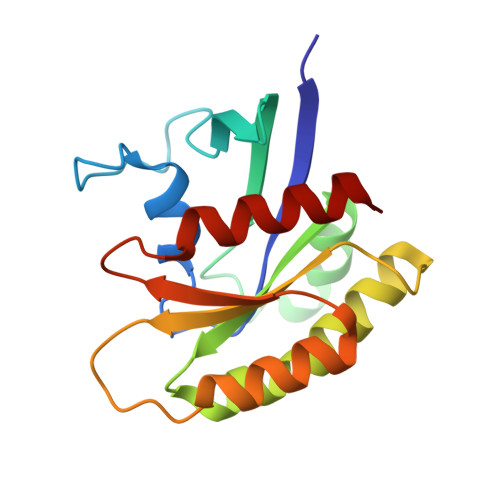
Discovery of Quinazolines That Activate SOS1-Mediated Nucleotide Exchange on RAS.
Abbott, J.R., Patel, P.A., Howes, J.E., Akan, D.T., Kennedy, J.P., Burns, M.C., Browning, C.F., Sun, Q., Rossanese, O.W., Phan, J., Waterson, A.G., Fesik, S.W.(2018) ACS Med Chem Lett 9: 941-946
- PubMed: 30258545 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsmedchemlett.8b00296
- Primary Citation Related Structures:
6CUO, 6CUP, 6CUR - PubMed Abstract:
Proteins in the RAS family are important regulators of cellular signaling and, when mutated, can drive cancer pathogenesis. Despite considerable effort over the last 30 years, RAS proteins have proven to be recalcitrant therapeutic targets. One approach for modulating RAS signaling is to target proteins that interact with RAS, such as the guanine nucleotide exchange factor (GEF) son of sevenless homologue 1 (SOS1). Here, we report hit-to-lead studies on quinazoline-containing compounds that bind to SOS1 and activate nucleotide exchange on RAS. Using structure-based design, we refined the substituents attached to the quinazoline nucleus and built in additional interactions not present in the initial HTS hit. Optimized compounds activate nucleotide exchange at single-digit micromolar concentrations in vitro. In HeLa cells, these quinazolines increase the levels of RAS-GTP and cause signaling changes in the mitogen-activated protein kinase/extracellular regulated kinase (MAPK/ERK) pathway.
- Department of Biochemistry and Department of Pharmacology, Vanderbilt University School of Medicine, Nashville, Tennessee 37232-0146, United States.
Organizational Affiliation: