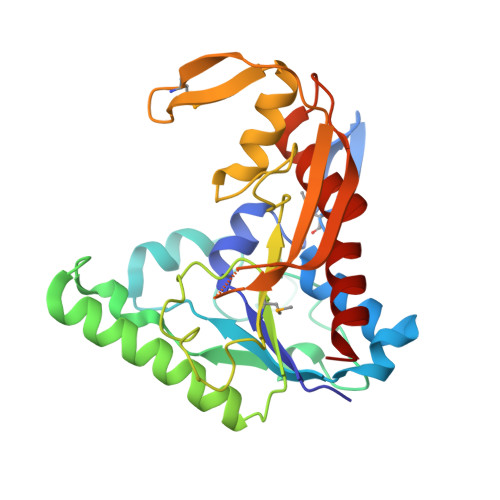
PvdF of pyoverdin biosynthesis is a structurally unique N10-formyltetrahydrofolate-dependent formyltransferase.
Kenjic, N., Hoag, M.R., Moraski, G.C., Caperelli, C.A., Moran, G.R., Lamb, A.L.(2019) Arch Biochem Biophys 664: 40-50
- PubMed: 30689984 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.abb.2019.01.028
- Primary Citation Related Structures:
6CUL - PubMed Abstract:
The hydroxyornithine transformylase from Pseudomonas aeruginosa is known by the gene name pvdF, and has been hypothesized to use N 10 -formyltetrahydrofolate (N 10 -fTHF) as a co-substrate formyl donor to convert N 5 -hydroxyornithine (OHOrn) to N 5 -formyl- N 5 -hydroxyornithine (fOHOrn). PvdF is in the biosynthetic pathway for pyoverdin biosynthesis, a siderophore generated under iron-limiting conditions that has been linked to virulence, quorum sensing and biofilm formation. The structure of PvdF was determined by X-ray crystallography to 2.3 Å, revealing a formyltransferase fold consistent with N 10 -formyltetrahydrofolate dependent enzymes, such as the glycinamide ribonucleotide transformylases, N-sugar transformylases and methionyl-tRNA transformylases. Whereas the core structure, including the catalytic triad, is conserved, PvdF has three insertions of 18 or more amino acids, which we hypothesize are key to binding the OHOrn substrate. Steady state kinetics revealed a non-hyperbolic rate curve, promoting the hypothesis that PvdF uses a random-sequential mechanism, and favors folate binding over OHOrn.
- Department of Molecular Biosciences, 1200 Sunnyside Ave, University of Kansas, Lawrence, KS, 66045, USA.
Organizational Affiliation: