Characteristic conformational changes on the distal and proximal surfaces of cytochrome P450 2D6 in response to substrate binding
Yang, Y.T., Fujita, F., Wang, P.F., Im, S.C., Pearl, N.M., Meagher, J., Stuckey, J., Waskell, L.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Cytochrome P450 2D6 | 479 | Homo sapiens | Mutation(s): 1 Gene Names: CYP2D6, CYP2DL1 EC: 1.14.14.1 (PDB Primary Data), 1.14.14 (UniProt) | 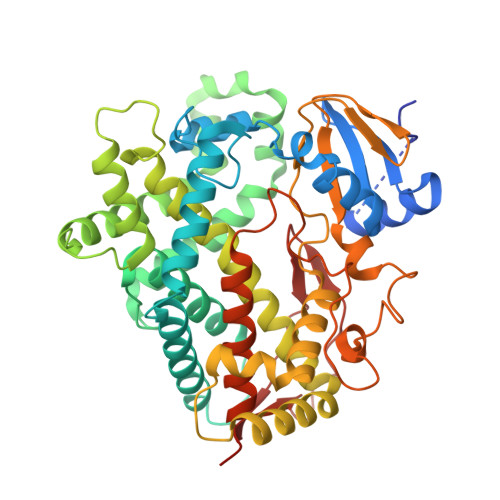 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P10635 GTEx: ENSG00000100197 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P10635 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 6 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| HEM Download:Ideal Coordinates CCD File | AA [auth D], E [auth A], L [auth B], T [auth C] | PROTOPORPHYRIN IX CONTAINING FE C34 H32 Fe N4 O4 KABFMIBPWCXCRK-RGGAHWMASA-L |  | ||
| 2CV Download:Ideal Coordinates CCD File | H [auth A], P [auth B], X [auth C] | HEGA-10 C18 H37 N O7 ITEIKACYSCODFV-ATLSCFEFSA-N |  | ||
| RTZ Download:Ideal Coordinates CCD File | BA [auth D] CA [auth D] F [auth A] G [auth A] M [auth B] | 10-{2-[(2R)-1-methylpiperidin-2-yl]ethyl}-2-(methylsulfanyl)-10H-phenothiazine C21 H26 N2 S2 KLBQZWRITKRQQV-MRXNPFEDSA-N |  | ||
| GOL Download:Ideal Coordinates CCD File | W [auth C] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| ZN Download:Ideal Coordinates CCD File | DA [auth D] I [auth A] J [auth A] K [auth A] Q [auth B] | ZINC ION Zn PTFCDOFLOPIGGS-UHFFFAOYSA-N |  | ||
| ACT Download:Ideal Coordinates CCD File | O [auth B] | ACETATE ION C2 H3 O2 QTBSBXVTEAMEQO-UHFFFAOYSA-M |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 57.738 | α = 90 |
| b = 191.999 | β = 90 |
| c = 250.46 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |
| PHASER | phasing |