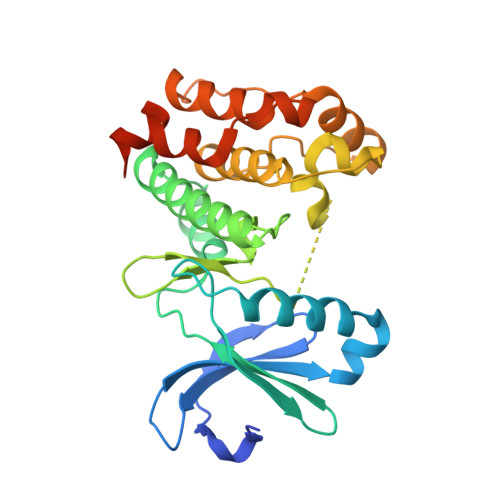
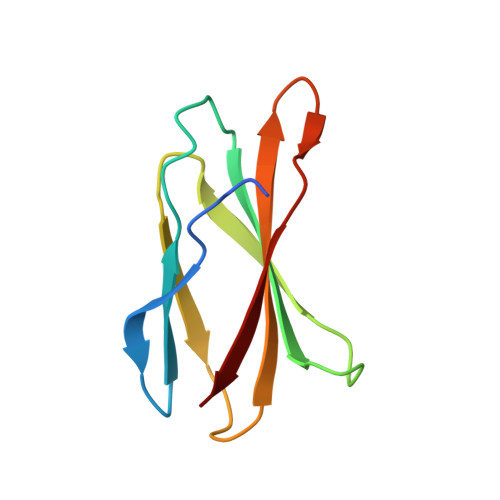
Dynamics of human protein kinase Aurora A linked to drug selectivity.
Pitsawong, W., Buosi, V., Otten, R., Agafonov, R.V., Zorba, A., Kern, N., Kutter, S., Kern, G., Padua, R.A., Meniche, X., Kern, D.(2018) Elife 7
- PubMed: 29901437 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.36656
- Primary Citation Related Structures:
6CPE, 6CPF, 6CPG - PubMed Abstract:
Protein kinases are major drug targets, but the development of highly-selective inhibitors has been challenging due to the similarity of their active sites. The observation of distinct structural states of the fully-conserved Asp-Phe-Gly (DFG) loop has put the concept of conformational selection for the DFG-state at the center of kinase drug discovery. Recently, it was shown that Gleevec selectivity for the Tyr-kinase Abl was instead rooted in conformational changes after drug binding. Here, we investigate whether protein dynamics after binding is a more general paradigm for drug selectivity by characterizing the binding of several approved drugs to the Ser/Thr-kinase Aurora A. Using a combination of biophysical techniques, we propose a universal drug-binding mechanism, that rationalizes selectivity, affinity and long on-target residence time for kinase inhibitors. These new concepts, where protein dynamics in the drug-bound state plays the crucial role, can be applied to inhibitor design of targets outside the kinome.
- Department of Biochemistry, Howard Hughes Medical Institute, Brandeis University, Waltham, United States.
Organizational Affiliation: