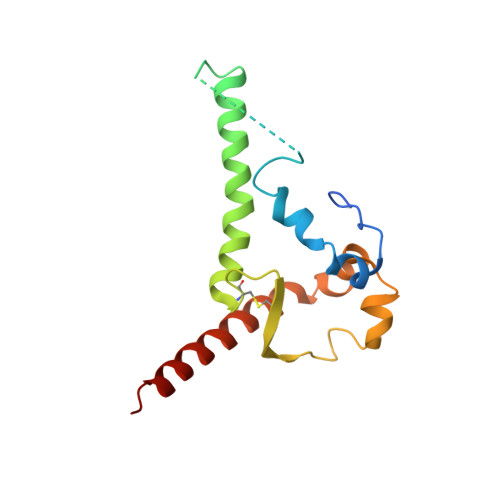
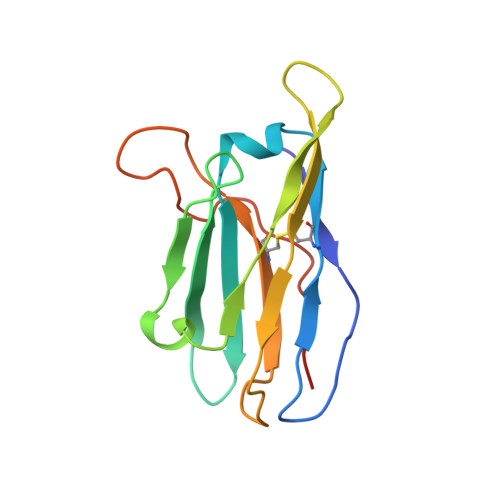
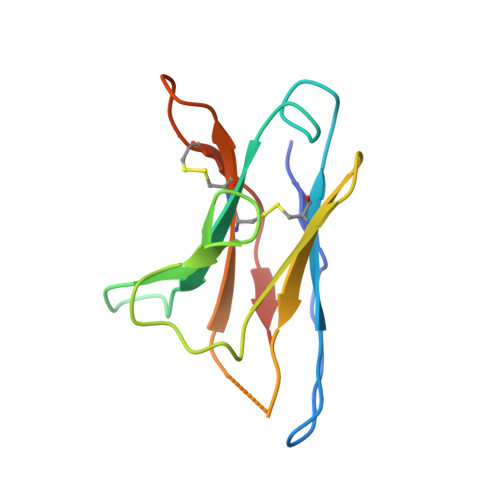
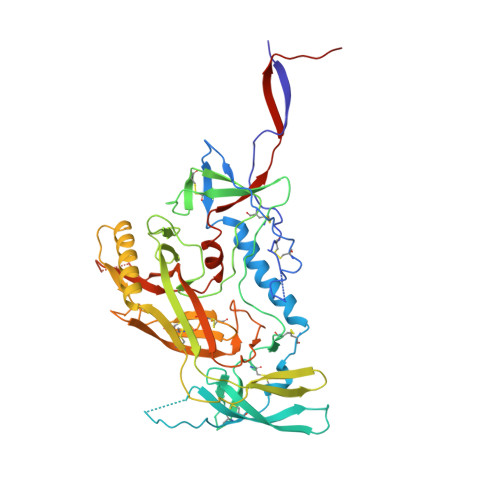
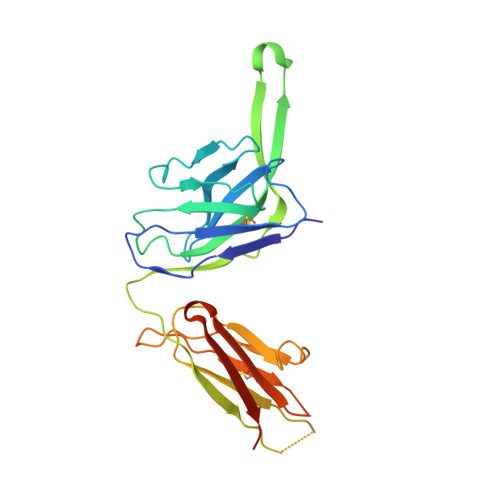
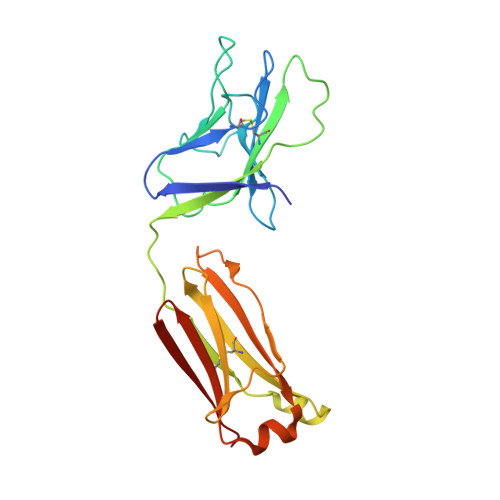
A Universal Approach to Optimize the Folding and Stability of Prefusion-Closed HIV-1 Envelope Trimers.
Rutten, L., Lai, Y.T., Blokland, S., Truan, D., Bisschop, I.J.M., Strokappe, N.M., Koornneef, A., van Manen, D., Chuang, G.Y., Farney, S.K., Schuitemaker, H., Kwong, P.D., Langedijk, J.P.M.(2018) Cell Rep 23: 584-595
- PubMed: 29642014 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2018.03.061
- Primary Citation Related Structures:
6CK9 - PubMed Abstract:
The heavily glycosylated native-like envelope (Env) trimer of HIV-1 is expected to have low immunogenicity, whereas misfolded forms are often highly immunogenic. High-quality correctly folded Envs may therefore be critical for developing a vaccine that induces broadly neutralizing antibodies. Moreover, the high variability of Env may require immunizations with multiple Envs. Here, we report a universal strategy that provides for correctly folded Env trimers of high quality and yield through a repair-and-stabilize approach. In the repair stage, we utilized a consensus strategy that substituted rare strain-specific residues with more prevalent ones. The stabilization stage involved structure-based design and experimental assessment confirmed by crystallographic feedback. Regions important for the refolding of Env were targeted for stabilization. Notably, the α9-helix and an intersubunit β sheet proved to be critical for trimer stability. Our approach provides a means to produce prefusion-closed Env trimers from diverse HIV-1 strains, a substantial advance for vaccine development.
- Janssen Vaccines & Prevention, Archimedesweg 4-6, Leiden 2333, the Netherlands.
Organizational Affiliation: