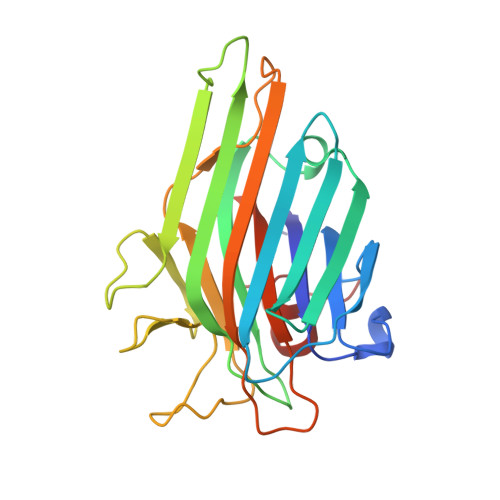
Crystal structure of DlyL, a mannose-specific lectin from Dioclea lasiophylla Mart. Ex Benth seeds that display cytotoxic effects against C6 glioma cells.
Leal, R.B., Pinto-Junior, V.R., Osterne, V.J.S., Wolin, I.A.V., Nascimento, A.P.M., Neco, A.H.B., Araripe, D.A., Welter, P.G., Neto, C.C., Correia, J.L.A., Rocha, C.R.C., Nascimento, K.S., Cavada, B.S.(2018) Int J Biol Macromol 114: 64-76
- PubMed: 29559315 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2018.03.080
- Primary Citation Related Structures:
6CJ9 - Universidade Federal de Santa Catarina (UFSC), Florianópolis, Santa Catarina, Brazil.
Organizational Affiliation: