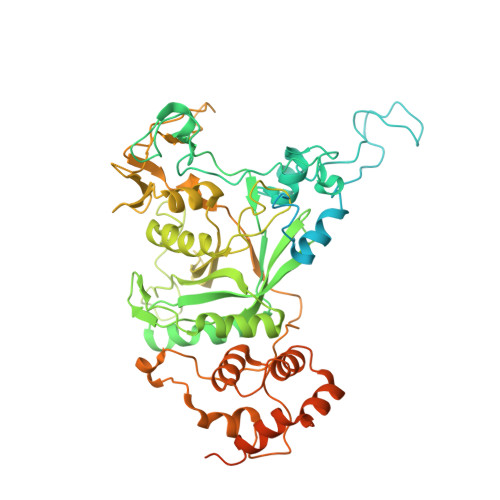
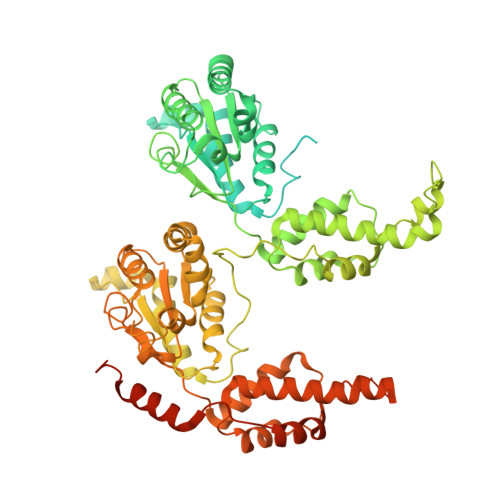
Structure of the Cdc48 ATPase with its ubiquitin-binding cofactor Ufd1-Npl4.
Bodnar, N.O., Kim, K.H., Ji, Z., Wales, T.E., Svetlov, V., Nudler, E., Engen, J.R., Walz, T., Rapoport, T.A.(2018) Nat Struct Mol Biol 25: 616-622
- PubMed: 29967539 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-018-0085-x
- Primary Citation Related Structures:
6CDD, 6CHS - PubMed Abstract:
Many polyubiquitinated proteins are extracted from membranes or complexes by the conserved ATPase Cdc48 (in yeast; p97 or VCP in mammals) before proteasomal degradation. Each Cdc48 hexamer contains two stacked ATPase rings (D1 and D2) and six N-terminal (N) domains. Cdc48 binds various cofactors, including the Ufd1-Npl4 heterodimer. Here, we report structures of the Cdc48-Ufd1-Npl4 complex from Chaetomium thermophilum. Npl4 interacts through its UBX-like domain with a Cdc48 N domain, and it uses two Zn 2+ -finger domains to anchor the enzymatically inactive Mpr1-Pad1 N-terminal (MPN) domain, homologous to domains found in several isopeptidases, to the top of the D1 ATPase ring. The MPN domain of Npl4 is located above Cdc48's central pore, a position similar to the MPN domain from deubiquitinase Rpn11 in the proteasome. Our results indicate that Npl4 is unique among Cdc48 cofactors and suggest a mechanism for binding and translocation of polyubiquitinated substrates into the ATPase.
- Howard Hughes Medical Institute and Department of Cell Biology, Harvard Medical School, Boston, MA, USA.
Organizational Affiliation: