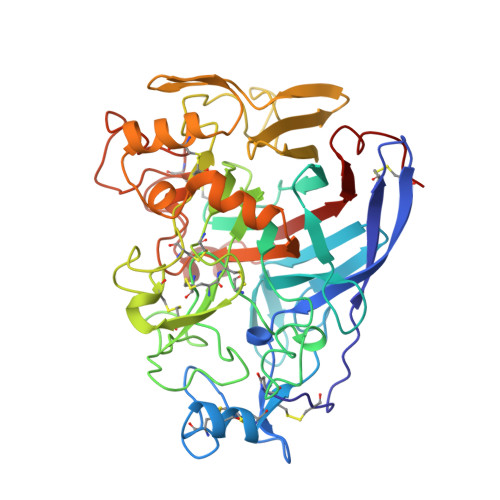
High-resolution crystal structures reveal how a cellulose chain is bound in the 50 A long tunnel of cellobiohydrolase I from Trichoderma reesei.
Divne, C., Stahlberg, J., Teeri, T.T., Jones, T.A.(1998) J Mol Biology 275: 309-325
- PubMed: 9466911 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1997.1437
- Primary Citation Related Structures:
5CEL, 6CEL, 7CEL - PubMed Abstract:
Detailed information has been obtained, by means of protein X-ray crystallography, on how a cellulose chain is bound in the cellulose-binding tunnel of cellobiohydrolase I (CBHI), the major cellulase in the hydrolysis of native, crystalline cellulose by the fungus Trichoderma reesei. Three high-resolution crystal structures of different catalytically deficient mutants of CBHI in complex with cellotetraose, cellopentaose and cellohexaose have been refined at 1.9, 1.7 and 1.9 A resolution, respectively. The observed binding of cellooligomers in the tunnel allowed unambiguous identification of ten well-defined subsites for glucosyl units that span a length of approximately 50 A. All bound oligomers have the same directionality and orientation, and the positions of the glucosyl units in each binding site agree remarkably well between the different complexes. The binding mode observed here corresponds to that expected during productive binding of a cellulose chain. The structures support the hypothesis that hydrolysis by CBHI proceeds from the reducing towards the non-reducing end of a cellulose chain, and they provide a structural explanation for the observed distribution of initial hydrolysis products.
- Department of Molecular Biology, Uppsala University, Sweden.
Organizational Affiliation: