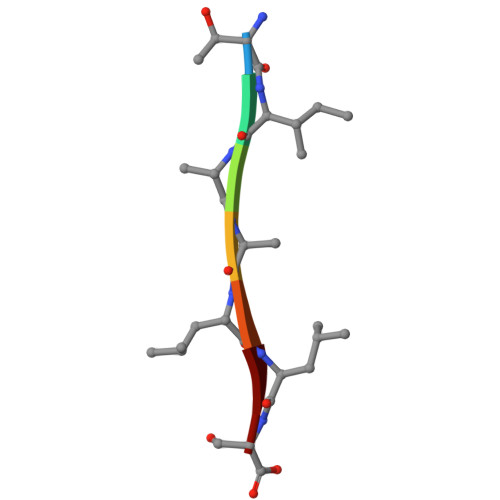
Crystal structures of amyloidogenic segments of human transthyretin.
Saelices, L., Sievers, S.A., Sawaya, M.R., Eisenberg, D.S.(2018) Protein Sci 27: 1295-1303
- PubMed: 29626847 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.3420
- Primary Citation Related Structures:
6C3F, 6C3G, 6C3S, 6C3T, 6C4O, 6C88 - PubMed Abstract:
Amyloid diseases are characterized by the deposition of proteins in the form of amyloid fibrils, in organs that eventually fail. The development of effective drug candidates follows from the understanding of the molecular processes that lead to protein aggregation. Here, we study amyloidogenic segments of transthyretin (TTR). TTR is a transporter of thyroxine and retinol in the blood and cerebrospinal fluid. When mutated and/or as a result of aging, TTR aggregates into amyloid fibrils that accumulate in organs such as the heart. Recently, we reported two amyloidogenic segments that drive amyloid aggregation. Here, we report the crystal structure of another six amyloidogenic segments of TTR. We found that the segments from the C-terminal region of TTR form in-register steric-zippers with highly-interdigitated, wet interfaces, whereas the β-strand B from the N-terminal region of TTR forms an out-of-register assembly, previously associated with oligomeric formation. Our results contribute fundamental information for understanding the mechanism of aggregation of TTR.
- Departments of Biological Chemistry and Chemistry and Biochemistry, Molecular Biology Institute, Box 951570, UCLA, Howard Hughes Medical Institute, UCLA-DOE Institute, Los Angeles, California, 90095-1570.
Organizational Affiliation: