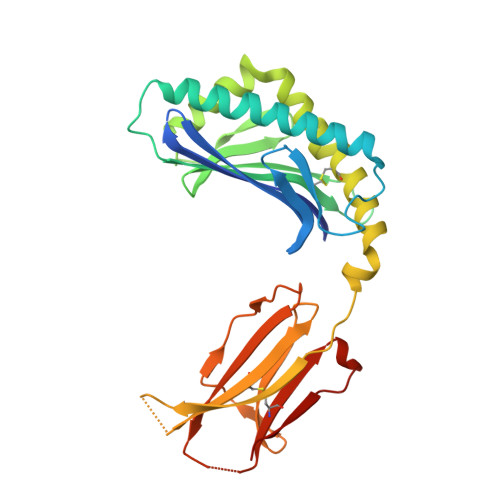
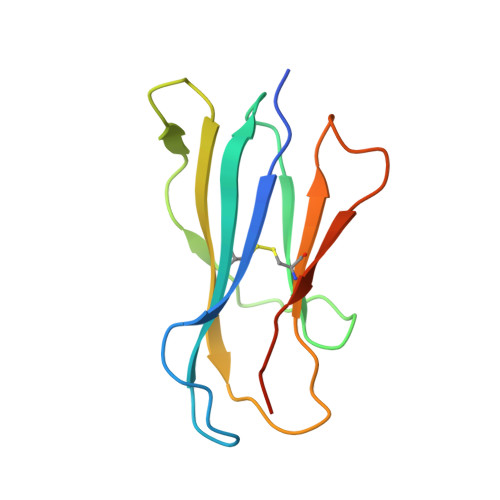
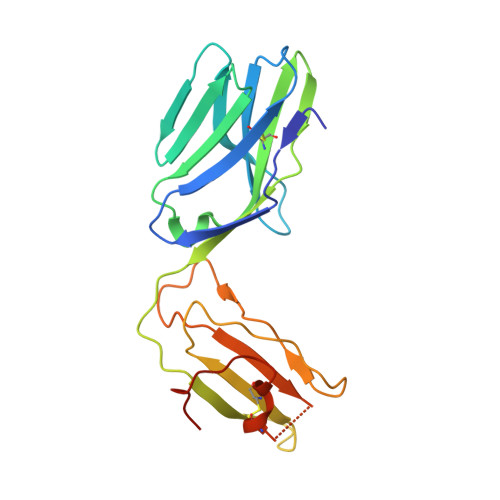
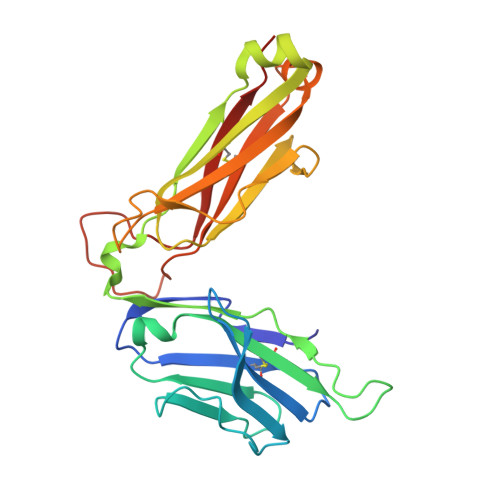
T cell autoreactivity directed toward CD1c itself rather than toward carried self lipids.
Wun, K.S., Reijneveld, J.F., Cheng, T.Y., Ladell, K., Uldrich, A.P., Le Nours, J., Miners, K.L., McLaren, J.E., Grant, E.J., Haigh, O.L., Watkins, T.S., Suliman, S., Iwany, S., Jimenez, J., Calderon, R., Tamara, K.L., Leon, S.R., Murray, M.B., Mayfield, J.A., Altman, J.D., Purcell, A.W., Miles, J.J., Godfrey, D.I., Gras, S., Price, D.A., Van Rhijn, I., Moody, D.B., Rossjohn, J.(2018) Nat Immunol 19: 397-406
- PubMed: 29531339 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41590-018-0065-7
- Primary Citation Related Structures:
6C09, 6C15 - PubMed Abstract:
The hallmark function of αβ T cell antigen receptors (TCRs) involves the highly specific co-recognition of a major histocompatibility complex molecule and its carried peptide. However, the molecular basis of the interactions of TCRs with the lipid antigen-presenting molecule CD1c is unknown. We identified frequent staining of human T cells with CD1c tetramers across numerous subjects. Whereas TCRs typically show high specificity for antigen, both tetramer binding and autoreactivity occurred with CD1c in complex with numerous, chemically diverse self lipids. Such extreme polyspecificity was attributable to binding of the TCR over the closed surface of CD1c, with the TCR covering the portal where lipids normally protrude. The TCR essentially failed to contact lipids because they were fully seated within CD1c. These data demonstrate the sequestration of lipids within CD1c as a mechanism of autoreactivity and point to small lipid size as a determinant of autoreactive T cell responses.
- Infection and Immunity Program and The Department of Biochemistry and Molecular Biology, Biomedicine Discovery Institute, Monash University, Clayton, Victoria, Australia.
Organizational Affiliation: