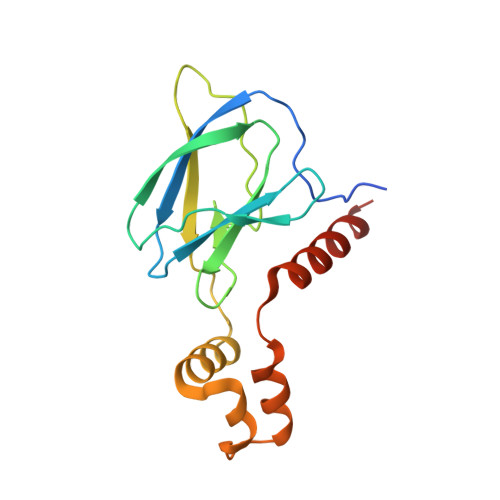
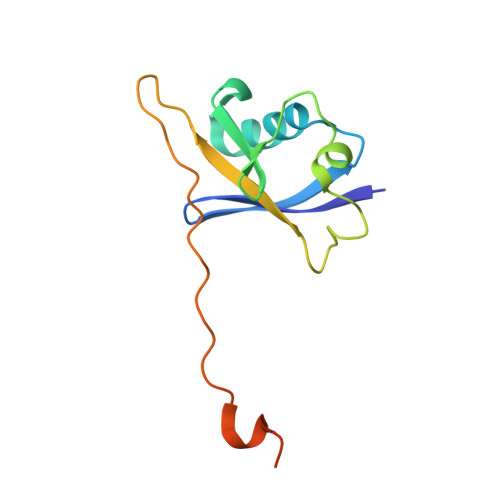
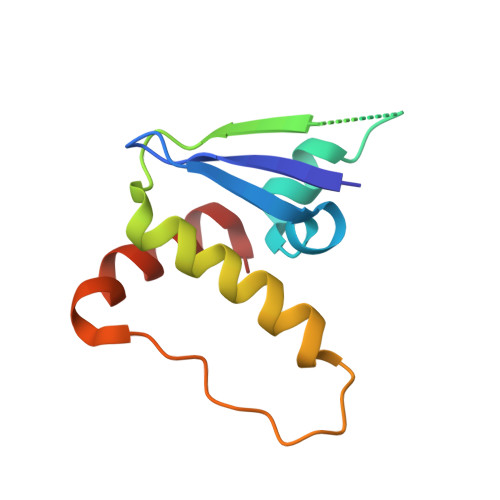
HIF-2 alpha-pVHL complex reveals broad genotype-phenotype correlations in HIF-2 alpha-driven disease.
Tarade, D., Robinson, C.M., Lee, J.E., Ohh, M.(2018) Nat Commun 9: 3359-3359
- PubMed: 30135421 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-05554-1
- Primary Citation Related Structures:
6BVB - PubMed Abstract:
It is definitively established that mutations in transcription factor HIF-2α are causative of both neuroendocrine tumors (class 1 disease) and polycythemia (class 2 disease). However, the molecular mechanism that underlies this emergent genotype-phenotype relationship has remained unclear. Here, we report the structure of HIF-2α peptide bound to pVHL-elongin B-elongin C (VBC) heterotrimeric complex, which shows topographical demarcation of class 1 and 2 mutations affecting residues predicted, and demonstrated via biophysical analyses, to differentially impact HIF-2α-pVHL interaction interface stability. Concordantly, biochemical experiments showed that class 1 mutations disrupt pVHL affinity to HIF-2α more adversely than class 2 mutations directly or indirectly via impeding PHD2-mediated hydroxylation. These findings suggest that neuroendocrine tumor pathogenesis requires a higher HIF-2α dose than polycythemia, which requires only a mild increase in HIF-2α activity. These biophysical data reveal a structural basis that underlies, and can be used to predict de novo, broad genotype-phenotype correlations in HIF-2α-driven disease.
- Department of Laboratory Medicine & Pathobiology, University of Toronto, 1 King's College Circle, Toronto, ON, M5S 1A8, Canada.
Organizational Affiliation: