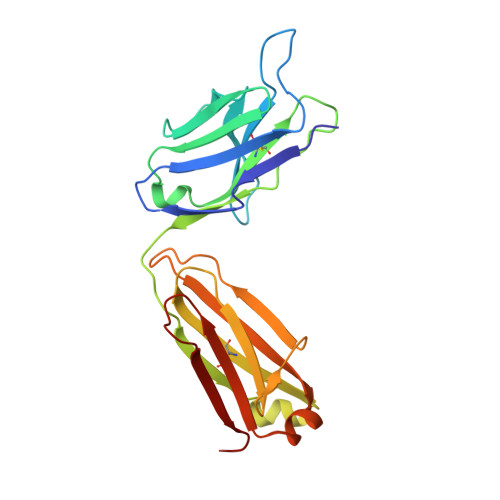
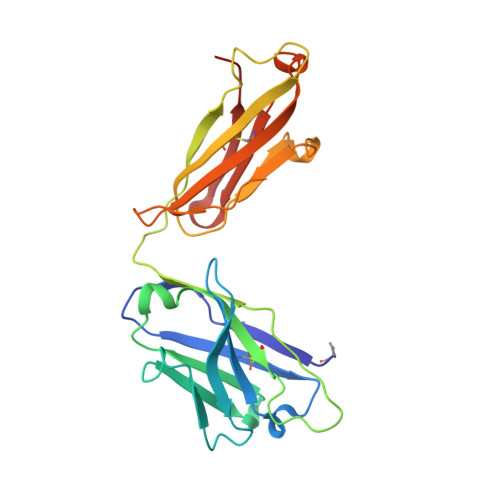
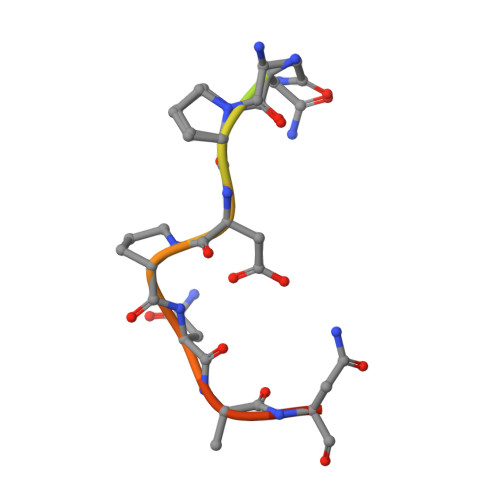
A public antibody lineage that potently inhibits malaria infection through dual binding to the circumsporozoite protein.
Tan, J., Sack, B.K., Oyen, D., Zenklusen, I., Piccoli, L., Barbieri, S., Foglierini, M., Fregni, C.S., Marcandalli, J., Jongo, S., Abdulla, S., Perez, L., Corradin, G., Varani, L., Sallusto, F., Sim, B.K.L., Hoffman, S.L., Kappe, S.H.I., Daubenberger, C., Wilson, I.A., Lanzavecchia, A.(2018) Nat Med 24: 401-407
- PubMed: 29554084 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nm.4513
- Primary Citation Related Structures:
6BQB - PubMed Abstract:
Immunization with attenuated Plasmodium falciparum sporozoites (PfSPZs) has been shown to be protective against malaria, but the features of the antibody response induced by this treatment remain unclear. To investigate this response in detail, we isolated IgM and IgG monoclonal antibodies from Tanzanian volunteers who were immunized with repeated injection of Sanaria PfSPZ Vaccine and who were found to be protected from controlled human malaria infection with infectious homologous PfSPZs. All isolated IgG monoclonal antibodies bound to P. falciparum circumsporozoite protein (PfCSP) and recognized distinct epitopes in its N terminus, NANP-repeat region, and C terminus. Strikingly, the most effective antibodies, as determined in a humanized mouse model, bound not only to the repeat region, but also to a minimal peptide at the PfCSP N-terminal junction that is not in the RTS,S vaccine. These dual-specific antibodies were isolated from different donors and were encoded by VH3-30 or VH3-33 alleles that encode tryptophan or arginine at position 52. Using structural and mutational data, we describe the elements required for germline recognition and affinity maturation. Our study provides potent neutralizing antibodies and relevant information for lineage-targeted vaccine design and immunization strategies.
- Institute for Research in Biomedicine, Università della Svizzera italiana, Bellinzona, Switzerland.
Organizational Affiliation: